Positive-strand RNA viruses (+ssRNA viruses) are a group of related viruses that have positive-sense, single-stranded genomes made of ribonucleic acid. The positive-sense genome can act as messenger RNA (mRNA) and can be directly translated into viral proteins by the host cell's ribosomes. Positive-strand RNA viruses encode an RNA-dependent RNA polymerase (RdRp) which is used during replication of the genome to synthesize a negative-sense antigenome that is then used as a template to create a new positive-sense viral genome.
| Positive-strand RNA virus | |
|---|---|
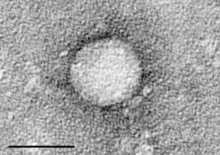
| |
| Hepatitis C virus | |
| Virus classification | |
| Group: | Group IV ((+)ssRNA)
|
| Kingdom: Phylum: Class | |
| Synonyms | |
| |
Positive-strand RNA viruses are divided between the phyla Kitrinoviricota, Lenarviricota, and Pisuviricota (specifically classes Pisoniviricetes and Stelpavirictes) all of which are in the kingdom Orthornavirae and realm Riboviria.[1] They are monophyletic and descended from a common RNA virus ancestor. In the Baltimore classification system, +ssRNA viruses belong to Group IV.[2]
Positive-sense RNA viruses include pathogens such as the Hepatitis C virus, West Nile virus, dengue virus, and the MERS, SARS, and SARS-CoV-2 coronaviruses,[3] as well as less clinically serious pathogens such as the coronaviruses and rhinoviruses that cause the common cold.[4][5][6]
Genome
editPositive-strand RNA virus genomes usually contain relatively few genes, usually between three and ten, including an RNA-dependent RNA polymerase.[4] Coronaviruses have the largest known RNA genomes, between 27 and 32 kilobases in length, and likely possess replication proofreading mechanisms in the form of an exoribonuclease within nonstructural protein nsp14.[7]
Replication
editPositive-strand RNA viruses have genetic material that can function both as a genome and as messenger RNA; it can be directly translated into protein in the host cell by host ribosomes.[8] The first proteins to be expressed after infection serve genome replication functions; they recruit the positive-strand viral genome to viral replication complexes formed in association with intracellular membranes. These complexes contain proteins of both viral and host cell origin, and may be associated with the membranes of a variety of organelles—often the rough endoplasmic reticulum,[9] but also including membranes derived from mitochondria, vacuoles, the Golgi apparatus, chloroplasts, peroxisomes, plasma membranes, autophagosomal membranes, and novel cytoplasmic compartments.[4]
The replication of the positive-sense RNA genome proceeds through double-stranded RNA intermediates, and the purpose of replication in these membranous invaginations may be the avoidance of cellular response to the presence of dsRNA. In many cases subgenomic RNAs are also created during replication.[8] After infection, the entirety of the host cell's translation machinery may be diverted to the production of viral proteins as a result of the very high affinity for ribosomes by the viral genome's internal ribosome entry site (IRES) elements; in some viruses, such as poliovirus and rhinoviruses, normal protein synthesis is further disrupted by viral proteases degrading components required to initiate translation of cellular mRNA.[6]
All positive-strand RNA virus genomes encode RNA-dependent RNA polymerase, a viral protein that synthesizes RNA from an RNA template. Host cell proteins recruited by +ssRNA viruses during replication include RNA-binding proteins, chaperone proteins, and membrane remodeling and lipid synthesis proteins, which collectively participate in exploiting the cell's secretory pathway for viral replication.[4]
Recombination
editNumerous positive-strand RNA viruses can undergo genetic recombination when at least two viral genomes are present in the same host cell.[10] The capability for recombination among +ssRNA virus pathogens of humans is common. RNA recombination appears to be a major driving force in determining genome architecture and the course of viral evolution among Picornaviridae (e.g. poliovirus).[11] In the Retroviridae (e.g. HIV), genome damage appears to be avoided during reverse transcription by strand switching, a form of recombination.[12][13][14] Recombination occurs in the Coronaviridae (e.g. SARS).[15] Recombination in RNA viruses appears to be an adaptation for coping with genome damage.[10] Recombination can also occur infrequently between +ssRNA viruses of the same species but of divergent lineages. The resulting recombinant viruses may sometimes cause an outbreak of infection in humans, as in the case of SARS and MERS.[15]
Positive-strand RNA viruses are common in plants. In tombusviruses and carmoviruses, RNA recombination occurs frequently during replication.[16] The ability of the RNA-dependent RNA polymerase of these viruses to switch RNA templates suggests a copy choice model of RNA recombination that may be an adaptive mechanism for coping with damage in the viral genome.[16] Other +ssRNA viruses of plants have also been reported to be capable of recombination, such as Brom mosaic bromovirus[17] and Sindbis virus.[18]
Classification
editPositive-strand RNA viruses are found in three phyla: Kitrinoviricota, Lenarviricota, and Pisuviricota, each of which are assigned to the kingdom Orthornavirae in the realm Riboviria. In the Baltimore classification system, which groups viruses together based on their manner of mRNA synthesis, +ssRNA viruses are group IV.[citation needed]
Kitrinoviricota
editThe first +ssRNA phylum is Kitrinoviricota. The phylum contains what have been referred to as the "alphavirus supergroup" and "flavivirus supergroup" along with various other short-genome viruses. Four classes in the phylum are recognized: Alsuviricetes, the alphavirus supergroup, which contains a large number of plant viruses and arthropod viruses; Flasuviricetes, which contains flaviviruses, Magsaviricetes, which contains nodaviruses and sinhaliviruses; and Tolucaviricetes, which primarily contains plant viruses.[19][20]
Lenarviricota
editLenarviricota is the second +ssRNA phylum. It contains the class Leviviricetes, which infect prokaryotes, and the apparent descendants of leviviruses, which infect eukaryotes. The phylum is divided into four classes: Leviviricetes, which contains leviviruses and their relatives, Amabiliviricetes, which contains narnaviruses and their relatives, Howeltoviricetes, which contains mitoviruses and their relatives, and Miaviricetes, which contains botourmiaviruses and their relatives. Based on phylogenetic analysis of RdRp, all other RNA viruses are considered to comprise a sister clade in relation to Lenarviricota.[19][20]
Pisuviricota
editThe third phylum that contains +ssRNA viruses is Pisuviricota, which has been informally called the "picornavirus supergroup". The phylum contains a large assemblage of eukaryotic viruses known to infect animals, plants, fungi, and protists. The phylum contains three classes, two of which contain only +ssRNA viruses: Pisoniviricetes, which contains nidoviruses, picornaviruses, and sobeliviruses, and Stelpaviricetes, which contains potyviruses and astroviruses. The third class is Duplopiviricetes, whose members are double-stranded RNA viruses that are descended from +ssRNA viruses.[19][20]
See also
editReferences
edit- ^ "Current ICTV Taxonomy Release | ICTV". ictv.global. Retrieved 3 April 2023.
- ^ Baltimore D (September 1971). "Expression of animal virus genomes". Bacteriological Reviews. 35 (3): 235–41. doi:10.1128/MMBR.35.3.235-241.1971. PMC 378387. PMID 4329869.
- ^ Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, Song H, Huang B, Zhu N, Bi Y, Ma X, Zhan F, Wang L, Hu T, Zhou H, Hu Z, Zhou W, Zhao L, Chen J, Meng Y, Wang J, Lin Y, Yuan J, Xie Z, Ma J, Liu WJ, Wang D, Xu W, Holmes EC, Gao GF, Wu G, Chen W, Shi W, Tan W (February 2020). "Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding". Lancet. 395 (10224): 565–574. doi:10.1016/S0140-6736(20)30251-8. PMC 7159086. PMID 32007145.
- ^ a b c d Nagy PD, Pogany J (December 2011). "The dependence of viral RNA replication on co-opted host factors". Nature Reviews. Microbiology. 10 (2): 137–149. doi:10.1038/nrmicro2692. PMC 7097227. PMID 22183253.
- ^ Ahlquist P, Noueiry AO, Lee WM, Kushner DB, Dye BT (August 2003). "Host factors in positive-strand RNA virus genome replication". Journal of Virology. 77 (15): 8181–8186. doi:10.1128/JVI.77.15.8181-8186.2003. PMC 165243. PMID 12857886.
- ^ a b Modrow S, Falke D, Truyen U, Schätzl H (2013). "Viruses with Single-Stranded, Positive-Sense RNA Genomes". Molecular Virology. Berlin, Heidelberg: Springer. pp. 185–349. doi:10.1007/978-3-642-20718-1_14. ISBN 978-3-642-20718-1. S2CID 82608215.
- ^ Smith EC, Denison MR (5 December 2013). "Coronaviruses as DNA wannabes: a new model for the regulation of RNA virus replication fidelity". PLOS Pathogens. 9 (12): e1003760. doi:10.1371/journal.ppat.1003760. PMC 3857799. PMID 24348241.
- ^ a b "Positive stranded RNA virus replication". ViralZone. Retrieved 8 September 2016.
- ^ Andronov L, Han M, Zhu Y, Balaji A, Roy AR, Barentine AE, Patel P, Garhyan J, Qi LS, Moerner WE (May 2024). "Nanoscale cellular organization of viral RNA and proteins in SARS-CoV-2 replication organelles". Nature Communications. 15 (1): 4644. Bibcode:2024NatCo..15.4644A. doi:10.1038/s41467-024-48991-x. PMC 11143195. PMID 38821943.
- ^ a b Barr JN, Fearns R (June 2010). "How RNA viruses maintain their genome integrity". The Journal of General Virology. 91 (Pt 6): 1373–87. doi:10.1099/vir.0.020818-0. PMID 20335491.
- ^ Muslin C, Mac Kain A, Bessaud M, Blondel B, Delpeyroux F (September 2019). "Recombination in Enteroviruses, a Multi-Step Modular Evolutionary Process". Viruses. 11 (9): 859. doi:10.3390/v11090859. PMC 6784155. PMID 31540135.
- ^ Hu WS, Temin HM (November 1990). "Retroviral recombination and reverse transcription". Science. 250 (4985): 1227–33. Bibcode:1990Sci...250.1227H. doi:10.1126/science.1700865. PMID 1700865.
- ^ Rawson JM, Nikolaitchik OA, Keele BF, Pathak VK, Hu WS (November 2018). "Recombination is required for efficient HIV-1 replication and the maintenance of viral genome integrity". Nucleic Acids Research. 46 (20): 10535–10545. doi:10.1093/nar/gky910. PMC 6237782. PMID 30307534.
- ^ Bernstein H, Bernstein C, Michod RE (January 2018). "Sex in microbial pathogens". Infection, Genetics and Evolution. 57: 8–25. Bibcode:2018InfGE..57....8B. doi:10.1016/j.meegid.2017.10.024. PMID 29111273.
- ^ a b Su S, Wong G, Shi W, Liu J, Lai AC, Zhou J, et al. (June 2016). "Epidemiology, Genetic Recombination, and Pathogenesis of Coronaviruses". Trends in Microbiology. 24 (6): 490–502. doi:10.1016/j.tim.2016.03.003. PMC 7125511. PMID 27012512.
- ^ a b Cheng CP, Nagy PD (November 2003). "Mechanism of RNA recombination in carmo- and tombusviruses: evidence for template switching by the RNA-dependent RNA polymerase in vitro". Journal of Virology. 77 (22): 12033–47. doi:10.1128/jvi.77.22.12033-12047.2003. PMC 254248. PMID 14581540.
- ^ Kolondam B, Rao P, Sztuba-Solinska J, Weber PH, Dzianott A, Johns MA, Bujarski JJ (2015). "Co-infection with two strains of Brome mosaic bromovirus reveals common RNA recombination sites in different hosts". Virus Evolution. 1 (1): vev021. doi:10.1093/ve/vev021. PMC 5014487. PMID 27774290.
- ^ Weiss BG, Schlesinger S (August 1991). "Recombination between Sindbis virus RNAs". Journal of Virology. 65 (8): 4017–25. doi:10.1128/JVI.65.8.4017-4025.1991. PMC 248832. PMID 2072444.
- ^ a b c Koonin EV, Dolja VV, Krupovic M, Varsani A, Wolf YI, Yutin N, Zerbini M, Kuhn JH (18 October 2019). "Create a megataxonomic framework, filling all principal taxonomic ranks, for realm Riboviria" (docx). International Committee on Taxonomy of Viruses (ICTV). Retrieved 14 August 2020.
- ^ a b c Wolf YI, Kazlauskas D, Iranzo J, Lucia-Sanz A, Kuhn JH, Krupovic M, Dolja VV, Koonin EV (27 November 2018). "Origins and Evolution of the Global RNA Virome". mBio. 9 (6): e02329-18. doi:10.1128/mBio.02329-18. PMC 6282212. PMID 30482837.