In molecular biology, Proteinase K (EC 3.4.21.64, protease K, endopeptidase K, Tritirachium alkaline proteinase, Tritirachium album serine proteinase, Tritirachium album proteinase K) is a broad-spectrum serine protease.[2][3][4] The enzyme was discovered in 1974 in extracts of the fungus Parengyodontium album (formerly Engyodontium album or Tritirachium album).[5] Proteinase K is able to digest hair (keratin), hence, the name "Proteinase K". The predominant site of cleavage is the peptide bond adjacent to the carboxyl group of aliphatic and aromatic amino acids with blocked alpha amino groups. It is commonly used for its broad specificity. This enzyme belongs to Peptidase family S8 (subtilisin). The molecular weight of Proteinase K is 28,900 daltons (28.9 kDa).
| Proteinase K | |||||||||
|---|---|---|---|---|---|---|---|---|---|
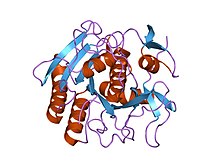 Structure of Proteinase K.[1] | |||||||||
| Identifiers | |||||||||
| EC no. | 3.4.21.64 | ||||||||
| CAS no. | 39450-01-6 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
| Proteinase K | |||||||
|---|---|---|---|---|---|---|---|
| Identifiers | |||||||
| Organism | |||||||
| Symbol | PROK | ||||||
| UniProt | P06873 | ||||||
| |||||||
Enzyme activity
editActivated by calcium, the enzyme digests proteins preferentially after hydrophobic amino acids (aliphatic, aromatic and other hydrophobic amino acids). Although calcium ions do not affect the enzyme activity, they do contribute to its stability. Proteins will be completely digested if the incubation time is long and the protease concentration high enough. Upon removal of the calcium ions, the stability of the enzyme is reduced, but the proteolytic activity remains.[6] Proteinase K has two binding sites for Ca2+, which are located close to the active center, but are not directly involved in the catalytic mechanism. The residual activity is sufficient to digest proteins, which usually contaminate nucleic acid preparations. Therefore, the digestion with Proteinase K for the purification of nucleic acids is usually performed in the presence of EDTA (inhibition of metal-ion dependent enzymes such as nucleases).
Proteinase K is also stable over a wide pH range (4–12), with a pH optimum of pH 8.0.[5] An elevation of the reaction temperature from 37 °C to 50–60 °C may increase the activity several times, like the addition of 0.5–1% sodium dodecyl sulfate (SDS) or Guanidinium chloride (3 M), Guanidinium thiocyanate (1 M) and urea (4 M) [disputed (for: no source cited for temperature) – discuss]. The above-mentioned conditions enhance proteinase K activity by making its substrate cleavage sites more accessible. Temperatures above 65 °C, trichloroacetic acid (TCA) or the serine protease-inhibitors AEBSF, PMSF or DFP inhibit the activity. Proteinase K will not be inhibited by Guanidinium chloride, Guanidinium thiocyanate, urea, Sarkosyl, Triton X-100, Tween 20, SDS, citrate, iodoacetic acid, EDTA or by other serine protease inhibitors like Nα-Tosyl-Lys Chloromethyl Ketone (TLCK) and Nα-Tosyl-Phe Chloromethyl Ketone (TPCK)[citation needed].
Protease K activity in commonly used buffers[7]
| Buffer (pH = 8.0, 50 °C, 1.25 μg/mL protease K, 15 min incubation) | Proteinase K activity (%) |
|---|---|
| 30 mM Tris·Cl | 100 |
| 30 mM Tris·Cl; 30 mM EDTA; 5% Tween 20; 0.5% Triton X-100; 800 mM GuHCl | 313 |
| 36 mM Tris·Cl; 36 mM EDTA; 5% Tween 20; 0.36% Triton X-100; 735 mM GuHCl | 301 |
| 10 mM Tris·Cl; 25 mM EDTA; 100 mM NaCl; 0.5% SDS | 128 |
| 10 mM Tris·Cl; 100 mM EDTA; 20 mM NaCl; 1% Sarkosyl | 74 |
| 10 mM Tris·Cl; 50 mM KCl; 1.5 mM MgCl2; 0.45% Tween 20; 0.5% Triton X-100 | 106 |
| 10 mM Tris·Cl; 100 mM EDTA; 0.5% SDS | 120 |
| 30 mM Tris·Cl; 10 mM EDTA; 1% SDS | 203 |
Applications
editProteinase K is commonly used in molecular biology to digest protein and remove contamination from preparations of nucleic acid. Addition of Proteinase K to nucleic acid preparations rapidly inactivates nucleases that might otherwise degrade the DNA or RNA during purification. It is highly suited to this application since the enzyme is active in the presence of chemicals that denature proteins, such as SDS and urea, chelating agents such as EDTA, sulfhydryl reagents, as well as trypsin or chymotrypsin inhibitors. Proteinase K is used for the destruction of proteins in cell lysates (tissue, cell culture cells) and for the release of nucleic acids, since it very effectively inactivates DNases and RNases. Some examples for applications: Proteinase K is very useful in the isolation of highly native, undamaged DNAs or RNAs, since most microbial or mammalian DNases and RNases are rapidly inactivated by the enzyme, particularly in the presence of 0.5–1% SDS.
The enzyme's activity towards native proteins is stimulated by denaturants such as SDS. In contrast, when measured using peptide substrates, denaturants inhibit the enzyme. The reason for this result is that the denaturing agents unfold the protein substrates and make them more accessible to the protease.[8]
Inhibitors
editProteinase K has two disulfide bonds,[9] but it exhibits higher proteolytic activity in the presence of reducing agents (e.g. 5 mM DTT),[10] suggesting that the presumed reduction of its own disulfide bonds does not lead to its irreversible inactivation. Proteinase K is inhibited by serine protease inhibitors such as phenylmethylsulfonyl fluoride (PMSF), diisopropylfluorophosphate (DFP), or 4-(2-aminoethyl)benzenesulfonyl fluoride (AEBSF). Proteinase K activity is unaffected by the sulfhydryl modifying reagents: para-chloromercuribenzoic acid (PCMB), N-alpha-tosyl-L-lysyl-chloromethyl-ketone (TLCK), or N-alpha-Tosyl-l-phenylalanine Chloromethyl Ketone (TPCK),[10] although presumably if these reagents were included alongside disulfide reducing reagents which exposed the typically-unavailable Proteinase K thiols, it may then become inhibited.
References
edit- ^ Betzel C, Singh TP, Visanji M, Peters K, Fittkau S, Saenger W, Wilson KS (July 1993). "Structure of the complex of proteinase K with a substrate analogue hexapeptide inhibitor at 2.2-A resolution". J. Biol. Chem. 268 (21): 15854–8. doi:10.1016/S0021-9258(18)82332-8. PMID 8340410.
- ^ Morihara K, Tsuzuki H (1975). "Specificity of proteinase K from Tritirachium album Limber for synthetic peptides". Agric. Biol. Chem. 39 (7): 1489–1492. doi:10.1271/bbb1961.39.1489.
- ^ Kraus E, Kiltz HH, Femfert UF (February 1976). "The specificity of proteinase K against oxidized insulin B chain". Hoppe-Seyler's Z. Physiol. Chem. 357 (2): 233–7. PMID 943367.
- ^ Jany KD, Lederer G, Mayer B (1986). "Amino acid sequence of proteinase K from the mold Tritirachium album Limber". FEBS Lett. 199 (2): 139–144. doi:10.1016/0014-5793(86)80467-7. S2CID 85577597.
- ^ a b Ebeling W, Hennrich N, Klockow M, Metz H, Orth HD, Lang H (August 1974). "Proteinase K from Tritirachium album Limber". Eur. J. Biochem. 47 (1): 91–7. doi:10.1111/j.1432-1033.1974.tb03671.x. PMID 4373242.
- ^ Müller A, Hinrichs W, Wolf WM, Saenger W (September 1994). "Crystal structure of calcium-free proteinase K at 1.5-A resolution". J. Biol. Chem. 269 (37): 23108–11. doi:10.2210/pdb2pkc/pdb. PMID 8083213.
- ^ Ag-Scientific. "Frequently asked questions about Proteinase K". Archived from the original on 17 October 2020. Retrieved 15 October 2020.
- ^ Hilz H, Wiegers U, Adamietz P (1975). "Stimulation of Proteinase K action by denaturing agents: application to the isolation of nucleic acids and the degradation of 'masked' proteins". European Journal of Biochemistry. 56 (1): 103–108. doi:10.1111/j.1432-1033.1975.tb02211.x. PMID 1236799.
- ^ "PROK - Proteinase K precursor - Parengyodontium album". Uniprot. 11 December 2019. Archived from the original on 19 November 2020. Retrieved 7 February 2020.
- ^ a b "Novagen Proteinase K protocol" (PDF). Sigma Aldrich. 11 December 2019. Archived (PDF) from the original on 6 June 2024. Retrieved 7 February 2020.
External links
edit- Proteinase K Worthington enzyme manual