The enzyme histidine decarboxylase (EC 4.1.1.22, HDC) is transcribed on chromosome 15, region q21.1-21.2, and catalyzes the decarboxylation of histidine to form histamine. In mammals, histamine is an important biogenic amine with regulatory roles in neurotransmission, gastric acid secretion and immune response.[1][2] Histidine decarboxylase is the sole member of the histamine synthesis pathway, producing histamine in a one-step reaction. Histamine cannot be generated by any other known enzyme.[citation needed] HDC is therefore the primary source of histamine in most mammals and eukaryotes. The enzyme employs a pyridoxal 5'-phosphate (PLP) cofactor, in similarity to many amino acid decarboxylases.[3][4] Eukaryotes, as well as gram-negative bacteria share a common HDC, while gram-positive bacteria employ an evolutionarily unrelated pyruvoyl-dependent HDC.[5] In humans, histidine decarboxylase is encoded by the HDC gene.[2][6]
| Histidine Decarboxylase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
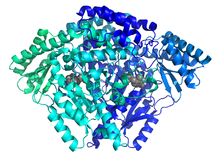 Cartoon depiction of C-truncated HDC dimer with PLP residing in active site. | |||||||||
| Identifiers | |||||||||
| EC no. | 4.1.1.22 | ||||||||
| CAS no. | 9024-61-7 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| Gene Ontology | AmiGO / QuickGO | ||||||||
| |||||||||
Structure
editHistidine decarboxylase is a group II pyridoxal-dependent decarboxylase, along with aromatic-L-amino-acid decarboxylase, and tyrosine decarboxylase. HDC is expressed as a 74 kDa polypeptide which is not enzymatically functional.[7][8] Only after post-translational processing does the enzyme become active. This processing consists of truncating much of the protein's C-terminal chain, reducing the peptide molecular weight to 54 kDa.
Histidine decarboxylase exists as a homodimer, with several amino acids from the respective opposing chain stabilizing the HDC active site. In HDC's resting state, PLP is covalently bound in a Schiff base to lysine 305, and stabilized by several hydrogen bonds to nearby amino acids aspartate 273, serine 151 and the opposing chain's serine 354.[7] HDC contains several regions that are sequentially and structurally similar to those in a number of other pyridoxal-dependent decarboxylases.[9] This is particularly evident in the vicinity of the active site lysine 305.[10]
Mechanism
editHDC decarboxylates histidine through the use of a PLP cofactor initially bound in a Schiff base to lysine 305.[11] Histidine initiates the reaction by displacing lysine 305 and forming an aldimine with PLP. Then, histidine's carboxyl group leaves the substrate, forming carbon dioxide. This is the rate-limiting step of the all process, requiring an activation energy of 17.6 kcal/mol [12] and fitting the experimental turnover of 1.73 .[13] After the decarboxylation takes place, the PLP intermediate is protonated by tyrosine 334 from the second subunit. The protonation is mediated by a water molecule and it is very fast and also very exergonic.[12] Finally, PLP re-forms its original Schiff base at lysine 305, and histamine is released. This mechanism is very similar to those employed by other pyridoxal-dependent decarboxylases. In particular, the aldimine intermediate is a common feature of all known PLP-dependent decarboxylases.[14] HDC is highly specific for its histidine substrate.[15]
Biological relevance
editHistidine decarboxylase is the primary biological source of histamine. Histamine is an important biogenic amine that moderates numerous physiologic processes. There are four different histamine receptors, H1, H2, H3, and H4,[16] each of which carries a different biological significance. H1 modulates several functions of the central and peripheral nervous system, including circadian rhythm, body temperature and appetite.[17] H2 activation results in gastric acid secretion and smooth muscle relaxation.[18][19] H3 controls histamine turnover by feedback inhibition of histamine synthesis and release.[20] Finally, H4 plays roles in mast cell chemotaxis and cytokine production.[17]
In humans, HDC is primarily expressed in mast cells and basophil granulocytes. Accordingly, these cells contain the body's highest concentrations of histamine granules. Non-mast cell histamine is also found in the brain, where it is used as a neurotransmitter.[21]
Inhibition
editHDC can be inhibited by α-fluoromethylhistidine and histidine methyl ester.[22][23]
Clinical significance
editAntihistamines are a class of medications designed to reduce unwanted effects of histamine in the body. Typical antihistamines block specific histamine receptors, depending on what physiological purpose they serve. For example, diphenhydramine (Benadryl™), targets and inhibits the H1 histamine receptor to relieve symptoms of allergic reactions.[24] Inhibitors of histidine decarboxylase can conceivably be used as atypical antihistamines. Tritoqualine, as well as various catechins, such as epigallocatechin-3-gallate, a major component of green tea, have been shown to target HDC and histamine-producing cells, reducing histamine levels and providing anti-inflammatory, anti-tumoral, and anti-angiogenic effects.[25]
Mutations in the gene for Histidine decarboxylase have been observed in one family with Tourette syndrome (TS) and are not thought to account for most cases of TS.[26]
See also
editReferences
edit- ^ Epps HM (1945). "Studies on bacterial amino-acid decarboxylases: 4. l(-)-histidine decarboxylase from Cl. welchii Type A". The Biochemical Journal. 39 (1): 42–6. doi:10.1042/bj0390042. PMC 1258146. PMID 16747851.
- ^ a b "Entrez Gene: histidine decarboxylase".
- ^ Riley WD, Snell EE (October 1968). "Histidine decarboxylase of Lactobacillus 30a. IV. The presence of covalently bound pyruvate as the prosthetic group". Biochemistry. 7 (10): 3520–8. doi:10.1021/bi00850a029. PMID 5681461.
- ^ Rosenthaler J, Guirard BM, Chang GW, Snell EE (July 1965). "Purification and properties of histidine decarboxylase from Lactobacillus 30a". Proceedings of the National Academy of Sciences of the United States of America. 54 (1): 152–8. Bibcode:1965PNAS...54..152R. doi:10.1073/pnas.54.1.152. PMC 285813. PMID 5216347.
- ^ Kimura B, Takahashi H, Hokimoto S, Tanaka Y, Fujii T (August 2009). "Induction of the histidine decarboxylase genes of Photobacterium damselae subsp. damselae (formally P. histaminum) at low pH". Journal of Applied Microbiology. 107 (2): 485–97. doi:10.1111/j.1365-2672.2009.04223.x. PMID 19302297.
- ^ Bruneau G, Nguyen VC, Gros F, Bernheim A, Thibault J (November 1992). "Preparation of a rat brain histidine decarboxylase (HDC) cDNA probe by PCR and assignment of the human HDC gene to chromosome 15". Human Genetics. 90 (3): 235–8. doi:10.1007/bf00220068. PMID 1487235. S2CID 23444983.
- ^ a b c Komori H, Nitta Y, Ueno H, Higuchi Y (August 2012). "Structural study reveals that Ser-354 determines substrate specificity on human histidine decarboxylase". The Journal of Biological Chemistry. 287 (34): 29175–83. doi:10.1074/jbc.M112.381897. PMC 3436558. PMID 22767596.
- ^ Nitta, Yoko (2010). "Expression of recombinant human histidine decarboxylase with full length and C-terminal truncated forms in yeast and bacterial cells" (PDF). J. Biol. Macromol. 10.
- ^ Jackson, F. Rob (1990-10-01). "Prokaryotic and eukaryotic pyridoxal-dependent decarboxylases are homologous". Journal of Molecular Evolution. 31 (4): 325–329. Bibcode:1990JMolE..31..325J. doi:10.1007/BF02101126. ISSN 0022-2844. PMID 2124279. S2CID 9206252.
- ^ Sandmeier E, Hale TI, Christen P (May 1994). "Multiple evolutionary origin of pyridoxal-5'-phosphate-dependent amino acid decarboxylases". European Journal of Biochemistry. 221 (3): 997–1002. doi:10.1111/j.1432-1033.1994.tb18816.x. PMID 8181483.
- ^ a b Wu F, Yu J, Gehring H (March 2008). "Inhibitory and structural studies of novel coenzyme-substrate analogs of human histidine decarboxylase". FASEB Journal. 22 (3): 890–7. doi:10.1096/fj.07-9566com. PMID 17965265. S2CID 39078427.
- ^ a b Fernandes HS, Ramos MJ, Cerqueira NM (July 2017). "The Catalytic Mechanism of the Pyridoxal-5'-phosphate-Dependent Enzyme, Histidine Decarboxylase: A Computational Study". Chemistry: A European Journal. 23 (38): 9162–9173. doi:10.1002/chem.201701375. PMID 28613002.
- ^ Komori H, Nitta Y, Ueno H, Higuchi Y (August 2012). "Structural study reveals that Ser-354 determines substrate specificity on human histidine decarboxylase". The Journal of Biological Chemistry. 287 (34): 29175–83. doi:10.1074/jbc.m112.381897. PMC 3436558. PMID 22767596.
- ^ "Pyridoxal phosphate-dependent decarboxylase". InterPro.
- ^ Toney MD (January 2005). "Reaction specificity in pyridoxal phosphate enzymes". Archives of Biochemistry and Biophysics. Highlight issue on Enzyme Mechanisms. 433 (1): 279–87. doi:10.1016/j.abb.2004.09.037. PMID 15581583.
- ^ Jutel, M.; Akdis, M.; Akdis, C. A. (2009-11-13). "Histamine, histamine receptors and their role in immune pathology". Clinical & Experimental Allergy. 39 (12): 1786–1800. doi:10.1111/j.1365-2222.2009.03374.x. ISSN 0954-7894. PMID 20085595.
- ^ a b Panula P, Chazot PL, Cowart M, Gutzmer R, Leurs R, Liu WL, Stark H, Thurmond RL, Haas HL (July 2015). "International Union of Basic and Clinical Pharmacology. XCVIII. Histamine Receptors". Pharmacological Reviews. 67 (3): 601–55. doi:10.1124/pr.114.010249. PMC 4485016. PMID 26084539.
- ^ Canonica GW, Blaiss M (February 2011). "Antihistaminic, anti-inflammatory, and antiallergic properties of the nonsedating second-generation antihistamine desloratadine: a review of the evidence". The World Allergy Organization Journal. 4 (2): 47–53. doi:10.1097/WOX.0b013e3182093e19. PMC 3500039. PMID 23268457.
- ^ Hill, S.J. (1997). "Classification of Histamine Receptors". Pharmacological Reviews. 49: 253–278 – via ASPET.
- ^ West RE, Zweig A, Shih NY, Siegel MI, Egan RW, Clark MA (November 1990). "Identification of two H3-histamine receptor subtypes". Molecular Pharmacology. 38 (5): 610–3. PMID 2172771.
- ^ Blandina P, Munari L, Provensi G, Passani MB (2012-01-01). "Histamine neurons in the tuberomamillary nucleus: a whole center or distinct subpopulations?". Frontiers in Systems Neuroscience. 6: 33. doi:10.3389/fnsys.2012.00033. PMC 3343474. PMID 22586376.
- ^ August TF, Musson DG, Hwang SS, Duggan DE, Hooke KF, Roman IJ, Ferguson RJ, Bayne WF (August 1985). "Bioanalysis and disposition of alpha-fluoromethylhistidine, a new histidine decarboxylase inhibitor". Journal of Pharmaceutical Sciences. 74 (8): 871–5. doi:10.1002/jps.2600740814. PMID 4032273.
- ^ Lane RS, Manning JM, Snell EE (September 1976). "Histidine decarboxylase of Lactobacillus 30a: inactivation and active-site labeling by L-histidine methyl ester". Biochemistry. 15 (19): 4180–5. doi:10.1021/bi00664a008. PMID 963031.
- ^ "Diphenhydramine Hydrochloride". Drugs.com.
- ^ Melgarejo E, Medina MA, Sánchez-Jiménez F, Urdiales JL (September 2010). "Targeting of histamine producing cells by EGCG: a green dart against inflammation?". Journal of Physiology and Biochemistry. 66 (3): 265–70. doi:10.1007/s13105-010-0033-7. PMID 20652470. S2CID 24835261.
- ^ "Online Mendelian Inheritance in Man: histidine decarboxylase".
Further reading
edit- Need AC, Keefe RS, Ge D, Grossman I, Dickson S, McEvoy JP, Goldstein DB (July 2009). "Pharmacogenetics of antipsychotic response in the CATIE trial: a candidate gene analysis". European Journal of Human Genetics. 17 (7): 946–57. doi:10.1038/ejhg.2008.264. PMC 2986499. PMID 19156168.
- Masini E, Fabbroni V, Giannini L, Vannacci A, Messerini L, Perna F, Cortesini C, Cianchi F (April 2005). "Histamine and histidine decarboxylase up-regulation in colorectal cancer: correlation with tumor stage" (PDF). Inflammation Research. 54 (Suppl 1): S80–1. doi:10.1007/s00011-004-0437-3. hdl:2158/762726. PMID 15928846. S2CID 28682686.
- Li Z, Liu J, Tang F, Liu Y, Waldum HL, Cui G (December 2008). "Expression of non-mast cell histidine decarboxylase in tumor-associated microvessels in human esophageal squamous cell carcinomas". APMIS. 116 (12): 1034–42. doi:10.1111/j.1600-0463.2008.01048.x. PMID 19133005. S2CID 19980875.
- Szafranski K, Schindler S, Taudien S, Hiller M, Huse K, Jahn N, Schreiber S, Backofen R, Platzer M (2007). "Violating the splicing rules: TG dinucleotides function as alternative 3' splice sites in U2-dependent introns". Genome Biology. 8 (8): R154. doi:10.1186/gb-2007-8-8-r154. PMC 2374985. PMID 17672918.
- Ai W, Liu Y, Langlois M, Wang TC (March 2004). "Kruppel-like factor 4 (KLF4) represses histidine decarboxylase gene expression through an upstream Sp1 site and downstream gastrin responsive elements". The Journal of Biological Chemistry. 279 (10): 8684–93. doi:10.1074/jbc.M308278200. PMID 14670968.
- Raychowdhury R, Fleming JV, McLaughlin JT, Bulitta CJ, Wang TC (October 2002). "Identification and characterization of a third gastrin response element (GAS-RE3) in the human histidine decarboxylase gene promoter". Biochemical and Biophysical Research Communications. 297 (5): 1089–95. doi:10.1016/S0006-291X(02)02345-8. PMID 12372397.
- Kimura K, Wakamatsu A, Suzuki Y, Ota T, Nishikawa T, Yamashita R, Yamamoto J, Sekine M, Tsuritani K, Wakaguri H, Ishii S, Sugiyama T, Saito K, Isono Y, Irie R, Kushida N, Yoneyama T, Otsuka R, Kanda K, Yokoi T, Kondo H, Wagatsuma M, Murakawa K, Ishida S, Ishibashi T, Takahashi-Fujii A, Tanase T, Nagai K, Kikuchi H, Nakai K, Isogai T, Sugano S (January 2006). "Diversification of transcriptional modulation: large-scale identification and characterization of putative alternative promoters of human genes". Genome Research. 16 (1): 55–65. doi:10.1101/gr.4039406. PMC 1356129. PMID 16344560.
- Sköldberg F, Portela-Gomes GM, Grimelius L, Nilsson G, Perheentupa J, Betterle C, Husebye ES, Gustafsson J, Rönnblom A, Rorsman F, Kämpe O (April 2003). "Histidine decarboxylase, a pyridoxal phosphate-dependent enzyme, is an autoantigen of gastric enterochromaffin-like cells". The Journal of Clinical Endocrinology and Metabolism. 88 (4): 1445–52. doi:10.1210/jc.2002-021761. PMID 12679420.
- Brew O, Lakasing L, Sullivan M (2007). "Differential activity of histidine decarboxylase in normal and pre-eclamptic placentae". Placenta. 28 (5–6): 585–7. doi:10.1016/j.placenta.2006.05.003. PMID 16822545.
- Zhang F, Xiong DH, Wang W, Shen H, Xiao P, Yang F, Recker RR, Deng HW (October 2006). "HDC gene polymorphisms are associated with age at natural menopause in Caucasian women". Biochemical and Biophysical Research Communications. 348 (4): 1378–82. doi:10.1016/j.bbrc.2006.08.008. PMC 1803761. PMID 16919600.
- Tippens AS, Gruetter CA (June 2004). "Detection of histidine decarboxylase mRNA in human vascular smooth muscle and endothelial cells". Inflammation Research. 53 (6): 215–6. doi:10.1007/s00011-004-1252-6. PMID 15167966.
- Siezen CL, Bont L, Hodemaekers HM, Ermers MJ, Doornbos G, Van't Slot R, Wijmenga C, Houwelingen HC, Kimpen JL, Kimman TG, Hoebee B, Janssen R (April 2009). "Genetic susceptibility to respiratory syncytial virus bronchiolitis in preterm children is associated with airway remodeling genes and innate immune genes". The Pediatric Infectious Disease Journal. 28 (4): 333–5. doi:10.1097/INF.0b013e31818e2aa9. PMID 19258923. S2CID 25601837.
- Morgan TK, Montgomery K, Mason V, West RB, Wang L, van de Rijn M, Higgins JP (July 2006). "Upregulation of histidine decarboxylase expression in superficial cortical nephrons during pregnancy in mice and women". Kidney International. 70 (2): 306–14. doi:10.1038/sj.ki.5001553. PMID 16760908.
- Papadopoulou N, Kalogeromitros D, Staurianeas NG, Tiblalexi D, Theoharides TC (November 2005). "Corticotropin-releasing hormone receptor-1 and histidine decarboxylase expression in chronic urticaria". The Journal of Investigative Dermatology. 125 (5): 952–5. doi:10.1111/j.0022-202X.2005.23913.x. PMID 16297195.
- Janssen R, Bont L, Siezen CL, Hodemaekers HM, Ermers MJ, Doornbos G, van 't Slot R, Wijmenga C, Goeman JJ, Kimpen JL, van Houwelingen HC, Kimman TG, Hoebee B (September 2007). "Genetic susceptibility to respiratory syncytial virus bronchiolitis is predominantly associated with innate immune genes". The Journal of Infectious Diseases. 196 (6): 826–34. doi:10.1086/520886. PMID 17703412.
- Strausberg RL, Feingold EA, Grouse LH, Derge JG, Klausner RD, Collins FS, et al. (December 2002). "Generation and initial analysis of more than 15,000 full-length human and mouse cDNA sequences". Proceedings of the National Academy of Sciences of the United States of America. 99 (26): 16899–903. Bibcode:2002PNAS...9916899M. doi:10.1073/pnas.242603899. PMC 139241. PMID 12477932.
- Aichberger KJ, Mayerhofer M, Vales A, Krauth MT, Gleixner KV, Bilban M, Esterbauer H, Sonneck K, Florian S, Derdak S, Pickl WF, Agis H, Falus A, Sillaber C, Valent P (November 2006). "The CML-related oncoprotein BCR/ABL induces expression of histidine decarboxylase (HDC) and the synthesis of histamine in leukemic cells". Blood. 108 (10): 3538–47. doi:10.1182/blood-2005-12-028456. PMID 16849647.
- Lee JK, Kim HT, Cho SM, Kim KH, Jin HJ, Ryu GM, Oh B, Park C, Kimm K, Jo SA, Jung SC, Kim S, In SM, Lee JE, Jo I (2003). "Characterization of 458 single nucleotide polymorphisms of disease candidate genes in the Korean population". Journal of Human Genetics. 48 (5): 213–6. doi:10.1007/s10038-003-0011-9. PMID 12768436.
- Jeong HJ, Moon PD, Kim SJ, Seo JU, Kang TH, Kim JJ, Kang IC, Um JY, Kim HM, Hong SH (April 2009). "Activation of hypoxia-inducible factor-1 regulates human histidine decarboxylase expression". Cellular and Molecular Life Sciences. 66 (7): 1309–19. doi:10.1007/s00018-009-9001-1. PMC 11131467. PMID 19266161. S2CID 23800803.
External links
edit- Histidine+Decarboxylase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
This article incorporates text from the United States National Library of Medicine, which is in the public domain.