Arabidopsis thaliana, the thale cress, mouse-ear cress or arabidopsis, is a small plant from the mustard family (Brassicaceae), native to Eurasia and Africa.[2][3][4][5][6][7] Commonly found along the shoulders of roads and in disturbed land, it is generally considered a weed.
| Arabidopsis thaliana | |
|---|---|

| |
| Scientific classification | |
| Kingdom: | Plantae |
| Clade: | Tracheophytes |
| Clade: | Angiosperms |
| Clade: | Eudicots |
| Clade: | Rosids |
| Order: | Brassicales |
| Family: | Brassicaceae |
| Genus: | Arabidopsis |
| Species: | A. thaliana
|
| Binomial name | |
| Arabidopsis thaliana | |
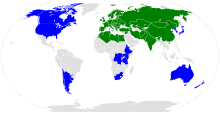
| |
The range of Arabidopsis thaliana.
| |
| Synonyms[1] | |
|
Arabis thaliana | |
A winter annual with a relatively short lifecycle, A. thaliana is a popular model organism in plant biology and genetics. For a complex multicellular eukaryote, A. thaliana has a relatively small genome of around 135 megabase pairs.[8] It was the first plant to have its genome sequenced, and is an important tool for understanding the molecular biology of many plant traits, including flower development and light sensing.[9]
Description
editArabidopsis thaliana is an annual (rarely biennial) plant, usually growing to 20–25 cm tall.[6] The leaves form a rosette at the base of the plant, with a few leaves also on the flowering stem. The basal leaves are green to slightly purplish in color, 1.5–5 cm long, and 2–10 mm broad, with an entire to coarsely serrated margin; the stem leaves are smaller and unstalked, usually with an entire margin. Leaves are covered with small, unicellular hairs called trichomes. The flowers are 3 mm in diameter, arranged in a corymb; their structure is that of the typical Brassicaceae. The fruit is a siliqua 5–20 mm long, containing 20–30 seeds.[10][11][12][13] Roots are simple in structure, with a single primary root that grows vertically downward, later producing smaller lateral roots. These roots form interactions with rhizosphere bacteria such as Bacillus megaterium.[14]
A. thaliana can complete its entire lifecycle in six weeks. The central stem that produces flowers grows after about 3 weeks, and the flowers naturally self-pollinate. In the lab, A. thaliana may be grown in Petri plates, pots, or hydroponics, under fluorescent lights or in a greenhouse.[15]
Taxonomy
editThe plant was first described in 1577 in the Harz Mountains by Johannes Thal (1542–1583), a physician from Nordhausen, Thüringen, Germany, who called it Pilosella siliquosa. In 1753, Carl Linnaeus renamed the plant Arabis thaliana in honor of Thal. In 1842, German botanist Gustav Heynhold erected the new genus Arabidopsis and placed the plant in that genus. The generic name, Arabidopsis, comes from Greek, meaning "resembling Arabis" (the genus in which Linnaeus had initially placed it).
Thousands of natural inbred accessions of A. thaliana have been collected from throughout its natural and introduced range.[16] These accessions exhibit considerable genetic and phenotypic variation, which can be used to study the adaptation of this species to different environments.[16]
Distribution and habitat
editA. thaliana is native to Europe, Asia, and Africa, and its geographic distribution is rather continuous from the Mediterranean to Scandinavia and Spain to Greece.[17] It also appears to be native in tropical alpine ecosystems in Africa and perhaps South Africa.[18][19] It has been introduced and naturalized worldwide,[20] including in North America around the 17th century.[21]
A. thaliana readily grows and often pioneers rocky, sandy, and calcareous soils. It is generally considered a weed, due to its widespread distribution in agricultural fields, roadsides, railway lines, waste ground, and other disturbed habitats,[20][22] but due to its limited competitive ability and small size, it is not categorized as a noxious weed.[23] Like most Brassicaceae species, A. thaliana is edible by humans in a salad or cooked, but it does not enjoy widespread use as a spring vegetable.[24]
Use as a model organism
editBotanists and biologists began to research A. thaliana in the early 1900s, and the first systematic description of mutants was done around 1945.[25] A. thaliana is now widely used for studying plant sciences, including genetics, evolution, population genetics, and plant development.[26][27][28] Although A. thaliana the plant has little direct significance for agriculture, A. thaliana the model organism has revolutionized our understanding of the genetic, cellular, and molecular biology of flowering plants.
The first mutant in A. thaliana was documented in 1873 by Alexander Braun, describing a double flower phenotype (the mutated gene was likely Agamous, cloned and characterized in 1990).[29] Friedrich Laibach (who had published the chromosome number in 1907) did not propose A. thaliana as a model organism, though, until 1943.[30] His student, Erna Reinholz, published her thesis on A. thaliana in 1945, describing the first collection of A. thaliana mutants that they generated using X-ray mutagenesis. Laibach continued his important contributions to A. thaliana research by collecting a large number of accessions (often questionably referred to as "ecotypes"). With the help of Albert Kranz, these were organised into a large collection of 750 natural accessions of A. thaliana from around the world.
In the 1950s and 1960s, John Langridge and George Rédei played an important role in establishing A. thaliana as a useful organism for biological laboratory experiments. Rédei wrote several scholarly reviews instrumental in introducing the model to the scientific community. The start of the A. thaliana research community dates to a newsletter called Arabidopsis Information Service,[31] established in 1964. The first International Arabidopsis Conference was held in 1965, in Göttingen, Germany.
In the 1980s, A. thaliana started to become widely used in plant research laboratories around the world. It was one of several candidates that included maize, petunia, and tobacco.[30] The latter two were attractive, since they were easily transformable with the then-current technologies, while maize was a well-established genetic model for plant biology. The breakthrough year for A. thaliana as a model plant was 1986, in which T-DNA-mediated transformation and the first cloned A. thaliana gene were described.[32][33]
Genomics
editNuclear genome
editDue to the small size of its genome, and because it is diploid, Arabidopsis thaliana is useful for genetic mapping and sequencing — with about 157 megabase pairs[36] and five chromosomes, A. thaliana has one of the smallest genomes among plants.[8] It was long thought to have the smallest genome of all flowering plants,[37] but that title is now considered to belong to plants in the genus Genlisea, order Lamiales, with Genlisea tuberosa, a carnivorous plant, showing a genome size of approximately 61 Mbp.[38] It was the first plant genome to be sequenced, completed in 2000 by the Arabidopsis Genome Initiative.[39] The most up-to-date version of the A. thaliana genome is maintained by the Arabidopsis Information Resource.[40]
The genome encodes ~27,600 protein-coding genes and about 6,500 non-coding genes.[41] However, the Uniprot database lists 39,342 proteins in their Arabidopsis reference proteome.[42] Among the 27,600 protein-coding genes 25,402 (91.8%) are now annotated with "meaningful" product names,[43] although a large fraction of these proteins is likely only poorly understood and only known in general terms (e.g. as "DNA-binding protein without known specificity"). Uniprot lists more than 3,000 proteins as "uncharacterized" as part of the reference proteome.
Chloroplast genome
editThe plastome of A. thaliana is a 154,478 base-pair-long DNA molecule,[34] a size typically encountered in most flowering plants (see the list of sequenced plastomes). It comprises 136 genes coding for small subunit ribosomal proteins (rps, in yellow: see figure), large subunit ribosomal proteins (rpl, orange), hypothetical chloroplast open reading frame proteins (ycf, lemon), proteins involved in photosynthetic reactions (green) or in other functions (red), ribosomal RNAs (rrn, blue), and transfer RNAs (trn, black).[35]
Mitochondrial genome
editThe mitochondrial genome of A. thaliana is 367,808 base pairs long and contains 57 genes.[44] There are many repeated regions in the Arabidopsis mitochondrial genome. The largest repeats recombine regularly and isomerize the genome.[45] Like most plant mitochondrial genomes, the Arabidopsis mitochondrial genome exists as a complex arrangement of overlapping branched and linear molecules in vivo.[46]
Genetics
editGenetic transformation of A. thaliana is routine, using Agrobacterium tumefaciens to transfer DNA into the plant genome. The current protocol, termed "floral dip", involves simply dipping flowers into a solution containing Agrobacterium carrying a plasmid of interest and a detergent.[47][48] This method avoids the need for tissue culture or plant regeneration.
The A. thaliana gene knockout collections are a unique resource for plant biology made possible by the availability of high-throughput transformation and funding for genomics resources. The site of T-DNA insertions has been determined for over 300,000 independent transgenic lines, with the information and seeds accessible through online T-DNA databases.[49] Through these collections, insertional mutants are available for most genes in A. thaliana.
Characterized accessions and mutant lines of A. thaliana serve as experimental material in laboratory studies. The most commonly used background lines are Ler (Landsberg erecta), and Col, or Columbia.[50] Other background lines less-often cited in the scientific literature are Ws, or Wassilewskija, C24, Cvi, or Cape Verde Islands, Nossen, etc. (see for ex.[51]) Sets of closely related accessions named Col-0, Col-1, etc., have been obtained and characterized; in general, mutant lines are available through stock centers, of which best-known are the Nottingham Arabidopsis Stock Center-NASC[50] and the Arabidopsis Biological Resource Center-ABRC in Ohio, USA.[52] The Col-0 accession was selected by Rédei from within a (nonirradiated) population of seeds designated 'Landsberg' which he received from Laibach.[53] Columbia (named for the location of Rédei's former institution, University of Missouri-Columbia) was the reference accession sequenced in the Arabidopsis Genome Initiative. The Later (Landsberg erecta) line was selected by Rédei (because of its short stature) from a Landsberg population he had mutagenized with X-rays. As the Ler collection of mutants is derived from this initial line, Ler-0 does not correspond to the Landsberg accessions, which designated La-0, La-1, etc.
Trichome formation is initiated by the GLABROUS1 protein. Knockouts of the corresponding gene lead to glabrous plants. This phenotype has already been used in gene editing experiments and might be of interest as visual marker for plant research to improve gene editing methods such as CRISPR/Cas9.[54][55]
Non-Mendelian inheritance controversy
editIn 2005, scientists at Purdue University proposed that A. thaliana possessed an alternative to previously known mechanisms of DNA repair, producing an unusual pattern of inheritance, but the phenomenon observed (reversion of mutant copies of the HOTHEAD gene to a wild-type state) was later suggested to be an artifact because the mutants show increased outcrossing due to organ fusion.[56][57][58]
Lifecycle
editThe plant's small size and rapid lifecycle are also advantageous for research. Having specialized as a spring ephemeral, it has been used to found several laboratory strains that take about 6 weeks from germination to mature seed. The small size of the plant is convenient for cultivation in a small space, and it produces many seeds. Further, the selfing nature of this plant assists genetic experiments. Also, as an individual plant can produce several thousand seeds, each of the above criteria leads to A. thaliana being valued as a genetic model organism.
Cellular biology
editArabidopsis is often the model for study of SNAREs in plants. This has shown SNAREs to be heavily involved in vesicle trafficking. Zheng et al. 1999 found an Arabidopsis SNARE called AtVTI1a is probably essential to Golgi-vacuole trafficking. This is still a wide open field and plant SNAREs' role in trafficking remains understudied.[59]
DNA repair
editThe DNA of plants is vulnerable to ultraviolet light, and DNA repair mechanisms have evolved to avoid or repair genome damage caused by UV. Kaiser et al.[60] showed that in A. thaliana cyclobutane pyrimidine dimers (CPDs) induced by UV light can be repaired by expression of CPD photolyase.
Germination in lunar regolith
editOn May 12, 2022, NASA announced that specimens of Arabidopsis thaliana had been successfully germinated and grown in samples of lunar regolith. While the plants successfully germinated and grew into seedlings, they were not as robust as specimens that had been grown in volcanic ash as a control group, although the experiments also found some variation in the plants grown in regolith based on the location the samples were taken from, as A. thaliana grown in regolith gathered during Apollo 12 & Apollo 17 were more robust than those grown in samples taken during Apollo 11.[61]
Development
editFlower development
editA. thaliana has been extensively studied as a model for flower development. The developing flower has four basic organs - sepals, petals, stamens, and carpels (which go on to form pistils). These organs are arranged in a series of whorls, four sepals on the outer whorl, followed by four petals inside this, six stamens, and a central carpel region. Homeotic mutations in A. thaliana result in the change of one organ to another—in the case of the agamous mutation, for example, stamens become petals and carpels are replaced with a new flower, resulting in a recursively repeated sepal-petal-petal pattern.
Observations of homeotic mutations led to the formulation of the ABC model of flower development by E. Coen and E. Meyerowitz.[62] According to this model, floral organ identity genes are divided into three classes - class A genes (which affect sepals and petals), class B genes (which affect petals and stamens), and class C genes (which affect stamens and carpels). These genes code for transcription factors that combine to cause tissue specification in their respective regions during development. Although developed through study of A. thaliana flowers, this model is generally applicable to other flowering plants.
Leaf development
editStudies of A. thaliana have provided considerable insights with regards to the genetics of leaf morphogenesis, particularly in dicotyledon-type plants.[63][64] Much of the understanding has come from analyzing mutants in leaf development, some of which were identified in the 1960s, but were not analysed with genetic and molecular techniques until the mid-1990s. A. thaliana leaves are well suited to studies of leaf development because they are relatively simple and stable.
Using A. thaliana, the genetics behind leaf shape development have become more clear and have been broken down into three stages: The initiation of the leaf primordium, the establishment of dorsiventrality, and the development of a marginal meristem. Leaf primordia are initiated by the suppression of the genes and proteins of class I KNOX family (such as SHOOT APICAL MERISTEMLESS). These class I KNOX proteins directly suppress gibberellin biosynthesis in the leaf primordium. Many genetic factors were found to be involved in the suppression of these class I KNOX genes in leaf primordia (such as ASYMMETRIC LEAVES1, BLADE-ON-PETIOLE1, SAWTOOTH1, etc.). Thus, with this suppression, the levels of gibberellin increase and leaf primordium initiate growth.
The establishment of leaf dorsiventrality is important since the dorsal (adaxial) surface of the leaf is different from the ventral (abaxial) surface.[65]
Microscopy
editA. thaliana is well suited for light microscopy analysis. Young seedlings on the whole, and their roots in particular, are relatively translucent. This, together with their small size, facilitates live cell imaging using both fluorescence and confocal laser scanning microscopy.[66] By wet-mounting seedlings in water or in culture media, plants may be imaged uninvasively, obviating the need for fixation and sectioning and allowing time-lapse measurements.[67] Fluorescent protein constructs can be introduced through transformation. The developmental stage of each cell can be inferred from its location in the plant or by using fluorescent protein markers, allowing detailed developmental analysis.
Physiology
editLight sensing, light emission, and circadian biology
editThe photoreceptors phytochromes A, B, C, D, and E mediate red light-based phototropic response. Understanding the function of these receptors has helped plant biologists understand the signaling cascades that regulate photoperiodism, germination, de-etiolation, and shade avoidance in plants. The genes FCA,[68] fy,[68] fpa,[68] LUMINIDEPENDENS (ld),[68] fly,[68] fve[68] and FLOWERING LOCUS C (FLC)[69][70] are involved in photoperiod triggering of flowering and vernalization. Specifically Lee et al 1994 find ld produces a homeodomain and Blazquez et al 2001 that fve produces a WD40 repeat.[68]
The UVR8 protein detects UV-B light and mediates the response to this DNA-damaging wavelength.
A. thaliana was used extensively in the study of the genetic basis of phototropism, chloroplast alignment, and stomal aperture and other blue light-influenced processes.[71] These traits respond to blue light, which is perceived by the phototropin light receptors. Arabidopsis has also been important in understanding the functions of another blue light receptor, cryptochrome, which is especially important for light entrainment to control the plants' circadian rhythms.[72] When the onset of darkness is unusually early, A. thaliana reduces its metabolism of starch by an amount that effectively requires division.[73]
Light responses were even found in roots, previously thought to be largely insensitive to light. While the gravitropic response of A. thaliana root organs is their predominant tropic response, specimens treated with mutagens and selected for the absence of gravitropic action showed negative phototropic response to blue or white light, and positive response to red light, indicating that the roots also show positive phototropism.[74]
In 2000, Dr. Janet Braam of Rice University genetically engineered A. thaliana to glow in the dark when touched. The effect was visible to ultrasensitive cameras.[75][better source needed]
Multiple efforts, including the Glowing Plant project, have sought to use A. thaliana to increase plant luminescence intensity towards commercially viable levels.
Thigmomorphogenesis (Touch response)
editIn 1990, Janet Braam and Ronald W. Davis determined that A. thaliana exhibits thigmomorphogenesis in response to wind, rain and touch.[76] Four or more touch induced genes in A. thaliana were found to be regulated by such stimuli.[76] In 2002, Massimo Pigliucci found that A. thaliana developed different patterns of branching in response to sustained exposure to wind, a display of phenotypic plasticity.[77]
On the Moon
editOn January 2, 2019, China's Chang'e-4 lander brought A. thaliana to the moon.[78] A small microcosm 'tin' in the lander contained A. thaliana, seeds of potatoes, and silkworm eggs. As plants would support the silkworms with oxygen, and the silkworms would in turn provide the plants with necessary carbon dioxide and nutrients through their waste,[79] researchers will evaluate whether plants successfully perform photosynthesis, and grow and bloom in the lunar environment.[78]
Secondary metabolites
editThalianin is an Arabidopsis root triterpene.[80] Potter et al., 2018 finds synthesis is induced by a combination of at least 2 facts, cell-specific transcription factors (TFs) and the accessibility of the chromatin.[80]
Plant–pathogen interactions
editUnderstanding how plants achieve resistance is important to protect the world's food production, and the agriculture industry. Many model systems have been developed to better understand interactions between plants and bacterial, fungal, oomycete, viral, and nematode pathogens. A. thaliana has been a powerful tool for the study of the subdiscipline of plant pathology, that is, the interaction between plants and disease-causing pathogens.
| Pathogen type | Example in A. thaliana |
|---|---|
| Bacteria | Pseudomonas syringae, Xanthomonas campestris |
| Fungi | Colletotrichum destructivum, Botrytis cinerea, Golovinomyces orontii |
| Oomycete | Hyaloperonospora arabidopsidis |
| Viral | Cauliflower mosaic virus (CaMV), tobacco mosaic virus (TMV) |
| Nematode | Meloidogyne incognita, Heterodera schachtii |
A schematic of PAMP-triggered immunity: recognition of flagellin by FLS2 (top left); effector-triggered immunity depicted through the recognition of avrRpt2 by RPS2 through RIN4 (top-right); microscopic view of callose deposition in an A. thaliana leaf (bottom left); an example of no hypersensitive response (HR) above, and HR in A. thaliana leaves below (bottom right)
on the roots of Arabidopsis thaliana
a) Overview of an A. thaliana root (primary root) with numerous root hairs, b) Biofilm-forming bacteria, c) Fungal or oomycete hyphae surrounding the root surface, d) Primary root densely covered by spores and protists, e, f) Protists, most likely belonging to the Bacillariophyceae class, g) Bacteria and bacterial filaments, h, i) Different bacterial individuals showing great varieties of shapes and morphological features[81]
The use of A. thaliana has led to many breakthroughs in the advancement of knowledge of how plants manifest plant disease resistance. The reason most plants are resistant to most pathogens is through nonhost resistance - not all pathogens will infect all plants. An example where A. thaliana was used to determine the genes responsible for nonhost resistance is Blumeria graminis, the causal agent of powdery mildew of grasses. A. thaliana mutants were developed using the mutagen ethyl methanesulfonate and screened to identify mutants with increased infection by B. graminis.[82][83][84] The mutants with higher infection rates are referred to as PEN mutants due to the ability of B. graminis to penetrate A. thaliana to begin the disease process. The PEN genes were later mapped to identify the genes responsible for nonhost resistance to B. graminis.
In general, when a plant is exposed to a pathogen, or nonpathogenic microbe, an initial response, known as PAMP-triggered immunity (PTI), occurs because the plant detects conserved motifs known as pathogen-associated molecular patterns (PAMPs).[85] These PAMPs are detected by specialized receptors in the host known as pattern recognition receptors (PRRs) on the plant cell surface.
The best-characterized PRR in A. thaliana is FLS2 (Flagellin-Sensing2), which recognizes bacterial flagellin,[86][87] a specialized organelle used by microorganisms for the purpose of motility, as well as the ligand flg22, which comprises the 22 amino acids recognized by FLS2. Discovery of FLS2 was facilitated by the identification of an A. thaliana ecotype, Ws-0, that was unable to detect flg22, leading to the identification of the gene encoding FLS2. FLS2 shows striking similarity to rice XA21, the first PRR isolated in 1995.[citation needed] Both flagellin and UV-C act similarly to increase homologous recombination in A. thaliana, as demonstrated by Molinier et al. 2006. Beyond this somatic effect, they found this to extend to subsequent generations of the plant.[88]
A second PRR, EF-Tu receptor (EFR), identified in A. thaliana, recognizes the bacterial EF-Tu protein, the prokaryotic elongation factor used in protein synthesis, as well as the laboratory-used ligand elf18.[89] Using Agrobacterium-mediated transformation, a technique that takes advantage of the natural process by which Agrobacterium transfers genes into host plants, the EFR gene was transformed into Nicotiana benthamiana, tobacco plant that does not recognize EF-Tu, thereby permitting recognition of bacterial EF-Tu[90] thereby confirming EFR as the receptor of EF-Tu.
Both FLS2 and EFR use similar signal transduction pathways to initiate PTI. A. thaliana has been instrumental in dissecting these pathways to better understand the regulation of immune responses, the most notable one being the mitogen-activated protein kinase (MAP kinase) cascade. Downstream responses of PTI include callose deposition, the oxidative burst, and transcription of defense-related genes.[91]
PTI is able to combat pathogens in a nonspecific manner. A stronger and more specific response in plants is that of effector-triggered immunity (ETI), which is dependent upon the recognition of pathogen effectors, proteins secreted by the pathogen that alter functions in the host, by plant resistance genes (R-genes), often described as a gene-for-gene relationship. This recognition may occur directly or indirectly via a guardee protein in a hypothesis known as the guard hypothesis. The first R-gene cloned in A. thaliana was RPS2 (resistance to Pseudomonas syringae 2), which is responsible for recognition of the effector avrRpt2.[92] The bacterial effector avrRpt2 is delivered into A. thaliana via the Type III secretion system of P. syringae pv. tomato strain DC3000. Recognition of avrRpt2 by RPS2 occurs via the guardee protein RIN4, which is cleaved.[clarification needed] Recognition of a pathogen effector leads to a dramatic immune response known as the hypersensitive response, in which the infected plant cells undergo cell death to prevent the spread of the pathogen.[93]
Systemic acquired resistance (SAR) is another example of resistance that is better understood in plants because of research done in A. thaliana. Benzothiadiazol (BTH), a salicylic acid (SA) analog, has been used historically as an antifungal compound in crop plants. BTH, as well as SA, has been shown to induce SAR in plants. The initiation of the SAR pathway was first demonstrated in A. thaliana in which increased SA levels are recognized by nonexpresser of PR genes 1 (NPR1)[94] due to redox change in the cytosol, resulting in the reduction of NPR1. NPR1, which usually exists in a multiplex (oligomeric) state, becomes monomeric (a single unit) upon reduction.[95] When NPR1 becomes monomeric, it translocates to the nucleus, where it interacts with many TGA transcription factors, and is able to induce pathogen-related genes such as PR1.[96] Another example of SAR would be the research done with transgenic tobacco plants, which express bacterial salicylate hydroxylase, nahG gene, requires the accumulation of SA for its expression[97]
Although not directly immunological, intracellular transport affects susceptibility by incorporating - or being tricked into incorporating - pathogen particles. For example, the Dynamin-related protein 2b/drp2b gene helps to move invaginated material into cells, with some mutants increasing PstDC3000 virulence even further.[98]
Evolutionary aspect of plant-pathogen resistance
editPlants are affected by multiple pathogens throughout their lifetimes. In response to the presence of pathogens, plants have evolved receptors on their cell surfaces to detect and respond to pathogens.[99] Arabidopsis thaliana is a model organism used to determine specific defense mechanisms of plant-pathogen resistance.[100] These plants have special receptors on their cell surfaces that allow for detection of pathogens and initiate mechanisms to inhibit pathogen growth.[100] They contain two receptors, FLS2 (bacterial flagellin receptor) and EF-Tu (bacterial EF-Tu protein), which use signal transduction pathways to initiate the disease response pathway.[100] The pathway leads to the recognition of the pathogen causing the infected cells to undergo cell death to stop the spread of the pathogen.[100] Plants with FLS2 and EF-Tu receptors have shown to have increased fitness in the population.[97] This has led to the belief that plant-pathogen resistance is an evolutionary mechanism that has built up over generations to respond to dynamic environments, such as increased predation and extreme temperatures.[97]
A. thaliana has also been used to study SAR.[101] This pathway uses benzothiadiazol, a chemical inducer, to induce transcription factors, mRNA, of SAR genes. This accumulation of transcription factors leads to inhibition of pathogen-related genes.[101]
Plant-pathogen interactions are important for an understanding of how plants have evolved to combat different types of pathogens that may affect them.[97] Variation in resistance of plants across populations is due to variation in environmental factors. Plants that have evolved resistance, whether it be the general variation or the SAR variation, have been able to live longer and hold off necrosis of their tissue (premature death of cells), which leads to better adaptation and fitness for populations that are in rapidly changing environments.[97] In the future, comparisons of the pathosystems of wild populations + their coevolved pathogens with wild-wild hybrids of known parentage may reveal new mechanisms of balancing selection. In life history theory we may find that A. thaliana maintains certain alleles due to pleitropy between plant-pathogen effects and other traits, as in livestock.[102]
Research in A. thaliana suggests that the immunity regulator protein family EDS1 in general co-evolved with the CCHELO family of nucleotide-binding–leucine-rich-repeat-receptors (NLRs). Xiao et al. 2005 have shown that the powdery mildew immunity mediated by A. thaliana's RPW8 (which has a CCHELO domain) is dependent on two members of this family: EDS1 itself and PAD4.[103]
RESISTANCE TO PSEUDOMONAS SYRINGAE 5/RPS5 is a disease resistance protein which guards AvrPphB SUSCEPTIBLE 1/PBS1. PBS1, as the name would suggest, is the target of AvrPphB, an effector produced by Pseudomonas syringae pv. phaseolicola.[104]
Other research
editOngoing research on A. thaliana is being performed on the International Space Station by the European Space Agency. The goals are to study the growth and reproduction of plants from seed to seed in microgravity.[105][106]
Plant-on-a-chip devices in which A. thaliana tissues can be cultured in semi-in vitro conditions have been described.[107] Use of these devices may aid understanding of pollen-tube guidance and the mechanism of sexual reproduction in A. thaliana.
Researchers at the University of Florida were able to grow the plant in lunar soil originating from the Sea of Tranquillity.[108]
Self-pollination
editA. thaliana is a predominantly self-pollinating plant with an outcrossing rate estimated at less than 0.3%.[109] An analysis of the genome-wide pattern of linkage disequilibrium suggested that self-pollination evolved roughly a million years ago or more.[110] Meioses that lead to self-pollination are unlikely to produce significant beneficial genetic variability. However, these meioses can provide the adaptive benefit of recombinational repair of DNA damages during formation of germ cells at each generation.[111] Such a benefit may have been sufficient to allow the long-term persistence of meioses even when followed by self-fertilization. A physical mechanism for self-pollination in A. thaliana is through pre-anthesis autogamy, such that fertilisation takes place largely before flower opening.
Databases and other resources
edit- TAIR and NASC:[50] curated sources for diverse genetic and molecular biology information, links to gene expression databases[112] etc.
- Arabidopsis Biological Resource Center (seed and DNA stocks)
- Nottingham Arabidopsis Stock Centre (seed and DNA stocks)
- Artade database
- AraDiv: a dataset of functional traits and leaf hyperspectral reflectance of Arabidopsis thaliana: see data repository
See also
edit- A. thaliana responses to salinity
- BZIP intron plant
- The Thaliana Bridge, installed in 2021 at Harlow Carr was inspired by the work of the botanical scientist Rachel Leech and represents the sequence of an Arabidopsis thaliana chromosome.[113]
- Novosphingobium arabidopsis, isolated from the rhizosphere of the plant
References
edit- ^ Warwick SI, Francis A, Al-Shehbaz IA (2016). "Brassicaceae species checklist and database". Species 2000 & ITIS Catalogue of Life (26 ed.). ISSN 2405-8858. Archived from the original on 9 December 2018. Retrieved 1 June 2016.
- ^ "Arabidopsis thaliana". Germplasm Resources Information Network. Agricultural Research Service, United States Department of Agriculture. Retrieved 11 December 2017.
- ^ Hoffmann MH (2002). "Biogeography of Arabidopsis thaliana (L.) Heynh. (Brassicaceae)". Journal of Biogeography. 29 (1): 125–134. Bibcode:2002JBiog..29..125H. doi:10.1046/j.1365-2699.2002.00647.x. S2CID 84959150.
- ^ Mitchell-Olds T (December 2001). "Arabidopsis thaliana and its wild relatives: a model system for ecology and evolution". Trends in Ecology & Evolution. 16 (12): 693–700. doi:10.1016/s0169-5347(01)02291-1.
- ^ Sharbel TF, Haubold B, Mitchell-Olds T (2000). "Genetic isolation by distance in Arabidopsis thaliana: biogeography and postglacial colonization of Europe". Molecular Ecology. 9 (12): 2109–2118. Bibcode:2000MolEc...9.2109S. doi:10.1046/j.1365-294x.2000.01122.x. PMID 11123622. S2CID 1788832.
- ^ a b Krämer U (March 2015). "Planting molecular functions in an ecological context with Arabidopsis thaliana". eLife. 4: –06100. doi:10.7554/eLife.06100. PMC 4373673. PMID 25807084.
- ^ Durvasula A, Fulgione A, Gutaker RM, Alacakaptan SI, Flood PJ, Neto C, Tsuchimatsu T, Burbano HA, Picó FX, Alonso-Blanco C, Hancock AM (May 2017). "Arabidopsis thaliana". Proceedings of the National Academy of Sciences of the United States of America. 114 (20): 5213–5218. doi:10.1073/pnas.1616736114. PMC 5441814. PMID 28473417.
- ^ a b "Genome Assembly". The Arabidopsis Information Resource. Archived from the original on 7 March 2021. Retrieved 29 March 2016.
- ^ "Nifty 50: ARABIDOPSIS -- A PLANT GENOME PROJECT". www.nsf.gov. Archived from the original on 3 January 2024. Retrieved 10 February 2023.
- ^ Flora of NW Europe: Arabidopsis thaliana Archived 8 December 2007 at the Wayback Machine
- ^ Blamey, M. & Grey-Wilson, C. (1989). Flora of Britain and Northern Europe. ISBN 0-340-40170-2
- ^ Flora of Pakistan: Arabidopsis thaliana Archived 18 June 2008 at the Wayback Machine
- ^ Flora of China: Arabidopsis thaliana Archived 5 October 2018 at the Wayback Machine
- ^ López-Bucio J, Campos-Cuevas JC, Hernández-Calderón E, Velásquez-Becerra C, Farías-Rodríguez R, Macías-Rodríguez LI, Valencia-Cantero E (February 2007). "Bacillus megaterium rhizobacteria promote growth and alter root-system architecture through an auxin- and ethylene-independent signaling mechanism in Arabidopsis thaliana". Molecular Plant-Microbe Interactions. 20 (2): 207–17. doi:10.1094/MPMI-20-2-0207. PMID 17313171.
- ^ Meinke DW, Cherry JM, Dean C, Rounsley SD, Koornneef M (October 1998). "Arabidopsis thaliana: a model plant for genome analysis". Science. 282 (5389): 662, 679–82. Bibcode:1998Sci...282..662M. CiteSeerX 10.1.1.462.4735. doi:10.1126/science.282.5389.662. PMID 9784120.
- ^ a b The 1001 Genomes Consortium (July 2016). "1,135 Genomes Reveal the Global Pattern of Polymorphism in Arabidopsis thaliana". Cell. 166 (2): 481–491. doi:10.1016/j.cell.2016.05.063. PMC 4949382. PMID 27293186.
{{cite journal}}: CS1 maint: numeric names: authors list (link) - ^ "Arabidopsis thaliana (L.) Heynh". www.gbif.org. Archived from the original on 1 June 2019. Retrieved 8 December 2018.
- ^ Hedberg, Olov (1957). "Afroalpine Vascular Plants: A Taxonomic Revision". Acta Universitatis Upsaliensis: Symbolae Botanicae Upsalienses. 15 (1): 1–144.
- ^ Fulgione A, Hancock AM (September 2018). "Archaic lineages broaden our view on the history of Arabidopsis thaliana". The New Phytologist. 219 (4): 1194–1198. doi:10.1111/nph.15244. hdl:21.11116/0000-0002-C3C7-1. PMID 29862511.
- ^ a b "Arabidopsis thaliana – Overview". Encyclopedia of Life. Archived from the original on 10 June 2016. Retrieved 31 May 2016.
- ^ Exposito-Alonso M, Becker C, Schuenemann VJ, Reiter E, Setzer C, Slovak R, Brachi B, Hagmann J, Grimm DG, Chen J, Busch W, Bergelson J, Ness RW, Krause J, Burbano HA, Weigel D (February 2018). "The rate and potential relevance of new mutations in a colonizing plant lineage". PLOS Genetics. 14 (2): e1007155. doi:10.1371/journal.pgen.1007155. PMC 5825158. PMID 29432421.
- ^ "Arabidopsis thaliana (thale cress)". Kew Gardens. Archived from the original on 28 February 2018. Retrieved 27 February 2018.
- ^ "State and Federal Noxious Weeds List | USDA PLANTS". plants.sc.egov.usda.gov. Archived from the original on 9 December 2018. Retrieved 8 December 2018.
- ^ "IRMNG". Encyclopedia of Life. Archived from the original on 1 April 2018.
- ^ [1] Archived 22 October 2016 at the Wayback Machine TAIR: About Arabidopsis
- ^ Rensink WA, Buell CR (June 2004). "Arabidopsis to rice. Applying knowledge from a weed to enhance our understanding of a crop species". Plant Physiology. 135 (2): 622–9. doi:10.1104/pp.104.040170. PMC 514098. PMID 15208410.
- ^ Coelho SM, Peters AF, Charrier B, Roze D, Destombe C, Valero M, Cock JM (December 2007). "Complex life cycles of multicellular eukaryotes: new approaches based on the use of model organisms" (PDF). Gene. 406 (1–2): 152–70. doi:10.1016/j.gene.2007.07.025. PMID 17870254. S2CID 24427325. Archived (PDF) from the original on 9 July 2021. Retrieved 29 June 2021.
- ^ Platt A, Horton M, Huang YS, Li Y, Anastasio AE, Mulyati NW, Agren J, Bossdorf O, Byers D, Donohue K, Dunning M, Holub EB, Hudson A, Le Corre V, Loudet O, Roux F, Warthmann N, Weigel D, Rivero L, Scholl R, Nordborg M, Bergelson J, Borevitz JO (February 2010). Novembre J (ed.). "The scale of population structure in Arabidopsis thaliana". PLOS Genetics. 6 (2): e1000843. doi:10.1371/journal.pgen.1000843. PMC 2820523. PMID 20169178.
- ^ Yanofsky MF, Ma H, Bowman JL, Drews GN, Feldmann KA, Meyerowitz EM (July 1990). "The protein encoded by the Arabidopsis homeotic gene agamous resembles transcription factors". Nature. 346 (6279): 35–9. Bibcode:1990Natur.346...35Y. doi:10.1038/346035a0. PMID 1973265. S2CID 4323431.
- ^ a b Meyerowitz EM (January 2001). "Prehistory and history of Arabidopsis research". Plant Physiology. 125 (1): 15–9. doi:10.1104/pp.125.1.15. PMC 1539315. PMID 11154286.
- ^ "About AIS". The Arabidopsis Information Resource. 8 November 2018. Archived from the original on 27 April 2021. Retrieved 25 April 2021.
- ^ Lloyd AM, Barnason AR, Rogers SG, Byrne MC, Fraley RT, Horsch RB (October 1986). "Transformation of Arabidopsis thaliana with Agrobacterium tumefaciens". Science. 234 (4775): 464–6. Bibcode:1986Sci...234..464L. doi:10.1126/science.234.4775.464. PMID 17792019. S2CID 22125701.
- ^ Chang C, Meyerowitz EM (March 1986). "Molecular cloning and DNA sequence of the Arabidopsis thaliana alcohol dehydrogenase gene". Proceedings of the National Academy of Sciences of the United States of America. 83 (5): 1408–12. Bibcode:1986PNAS...83.1408C. doi:10.1073/pnas.83.5.1408. PMC 323085. PMID 2937058.
- ^ a b "Arabidopsis thaliana chloroplast, complete genome — NCBI accession number NC_000932.1". National Center for Biotechnology Information. Archived from the original on 4 November 2018. Retrieved 4 November 2018.
- ^ a b Sato S, Nakamura Y, Kaneko T, Asamizu E, Tabata S (1999). "Complete structure of the chloroplast genome of Arabidopsis thaliana". DNA Research. 6 (5): 283–290. doi:10.1093/dnares/6.5.283. ISSN 1340-2838. PMID 10574454.
- ^ Bennett MD, Leitch IJ, Price HJ, Johnston JS (April 2003). "Comparisons with Caenorhabditis (approximately 100 Mb) and Drosophila (approximately 175 Mb) using flow cytometry show genome size in Arabidopsis to be approximately 157 Mb, thus approximately 25% larger than the Arabidopsis genome initiative estimate of approximately 125 Mb". Annals of Botany. 91 (5): 547–57. doi:10.1093/aob/mcg057. PMC 4242247. PMID 12646499.
- ^ (Leutwileret al., 1984). In our survey Arabidopsis ...
- ^ Fleischmann A, Michael TP, Rivadavia F, Sousa A, Wang W, Temsch EM, Greilhuber J, Müller KF, Heubl G (December 2014). "Evolution of genome size and chromosome number in the carnivorous plant genus Genlisea (Lentibulariaceae), with a new estimate of the minimum genome size in angiosperms". Annals of Botany. 114 (8): 1651–63. doi:10.1093/aob/mcu189. PMC 4649684. PMID 25274549.
- ^ The Arabidopsis Genome Initiative (December 2000). "Analysis of the genome sequence of the flowering plant Arabidopsis thaliana". Nature. 408 (6814): 796–815. Bibcode:2000Natur.408..796T. doi:10.1038/35048692. PMID 11130711.
- ^ "TAIR - Genome Annotation". Archived from the original on 14 October 2008. Retrieved 29 December 2008.
- ^ "Details - Arabidopsis_thaliana - Ensembl Genomes 63". ensembl.gramene.org. Archived from the original on 24 June 2021. Retrieved 15 June 2021.
- ^ "Arabidopsis thaliana (Mouse-ear cress)". www.uniprot.org. Archived from the original on 21 May 2021. Retrieved 15 June 2021.
- ^ Cheng, Chia-Yi; Krishnakumar, Vivek; Chan, Agnes P.; Thibaud-Nissen, Françoise; Schobel, Seth; Town, Christopher D. (2017). "Araport11: a complete reannotation of the Arabidopsis thaliana reference genome". The Plant Journal. 89 (4): 789–804. doi:10.1111/tpj.13415. ISSN 1365-313X. PMID 27862469. S2CID 12155857.
- ^ "Arabidopsis thaliana ecotype Col-0 mitochondrion, complete genome — NCBI accession number BK010421". National Center for Biotechnology Information. 10 October 2018. Archived from the original on 12 April 2019. Retrieved 10 April 2019.
- ^ Klein M, Eckert-Ossenkopp U, Schmiedeberg I, Brandt P, Unseld M, Brennicke A, Schuster W (1994). "Physical mapping of the mitochondrial genome of Arabidopsis thaliana by cosmid and YAC clones". Plant Journal. 6 (3): 447–455. doi:10.1046/j.1365-313X.1994.06030447.x. PMID 7920724.
- ^ Gualberto JM, Mileshina D, Wallet C, Niazi AK, Weber-Lotfi F, Dietrich A (2014). "The plant mitochondrial genome: dynamics and maintenance". Biochimie. 100: 107–120. doi:10.1016/j.biochi.2013.09.016. PMID 24075874.
- ^ Clough SJ, Bent AF (December 1998). "Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana". The Plant Journal. 16 (6): 735–43. doi:10.1046/j.1365-313x.1998.00343.x. PMID 10069079. S2CID 410286.
- ^ Zhang X, Henriques R, Lin SS, Niu QW, Chua NH (2006). "Agrobacterium-mediated transformation of Arabidopsis thaliana using the floral dip method". Nature Protocols. 1 (2): 641–6. doi:10.1038/nprot.2006.97. PMID 17406292. S2CID 6906570.
- ^ "T-DNA Express: Arabidopsis Gene Mapping Tool". signal.salk.edu. Archived from the original on 25 November 2009. Retrieved 19 October 2009.
- ^ a b c "Eurasian Arabidopsis Stock Centre (uNASC)". arabidopsis.info. Archived from the original on 12 December 2001. Retrieved 19 October 2009.
- ^ Magliano TM, Botto JF, Godoy AV, Symonds VV, Lloyd AM, Casal JJ (June 2005). "New Arabidopsis recombinant inbred lines (Landsberg erecta x Nossen) reveal natural variation in phytochrome-mediated responses". Plant Physiology. 138 (2): 1126–35. doi:10.1104/pp.104.059071. PMC 1150426. PMID 15908601.
- ^ "ABRC". abrc.osu.edu. Archived from the original on 25 February 2021. Retrieved 12 December 2020.
- ^ "NASC Collection Info". arabidopsis.info. Archived from the original on 19 July 2011. Retrieved 15 February 2011.
- ^ Hahn F, Mantegazza O, Greiner A, Hegemann P, Eisenhut M, Weber AP (2017). "Arabidopsis thaliana". Frontiers in Plant Science. 8: 39. doi:10.3389/fpls.2017.00039. PMC 5258748. PMID 28174584.
- ^ Hahn F, Eisenhut M, Mantegazza O, Weber AP (5 April 2018). "Arabidopsis With Cas9-Based Gene Targeting". Frontiers in Plant Science. 9: 424. doi:10.3389/fpls.2018.00424. PMC 5895730. PMID 29675030.
- ^ Lolle SJ, Victor JL, Young JM, Pruitt RE (March 2005). "Genome-wide non-mendelian inheritance of extra-genomic information in Arabidopsis". Nature. 434 (7032): 505–9. Bibcode:2005Natur.434..505L. doi:10.1038/nature03380. PMID 15785770. S2CID 1352368.Washington Post summary. Archived 18 November 2016 at the Wayback Machine
- ^ Peng P, Chan SW, Shah GA, Jacobsen SE (September 2006). "Plant genetics: increased outcrossing in hothead mutants". Nature. 443 (7110): E8, discussion E8–9. Bibcode:2006Natur.443E...8P. doi:10.1038/nature05251. PMID 17006468. S2CID 4420979.
- ^ Pennisi E (September 2006). "Genetics. Pollen contamination may explain controversial inheritance". Science. 313 (5795): 1864. doi:10.1126/science.313.5795.1864. PMID 17008492. S2CID 82215542.
- ^ Raikhel, Natasha V. (28 April 2017). "Firmly Planted, Always Moving". Annual Review of Plant Biology. 68 (1). Annual Reviews: 1–27. doi:10.1146/annurev-arplant-042916-040829. ISSN 1543-5008. PMID 27860488.
- ^ Kaiser G, Kleiner O, Beisswenger C, Batschauer A. Increased DNA repair in Arabidopsis plants overexpressing CPD photolyase. Planta. 2009 Aug;230(3):505-15. doi: 10.1007/s00425-009-0962-y. Epub 2009 Jun 12. PMID 19521716
- ^ Keeter, Bill (12 May 2022). "Scientists Grow Plants in Lunar Soil". NASA. Archived from the original on 14 May 2022. Retrieved 14 May 2022.
- ^ Coen ES, Meyerowitz EM (September 1991). "The war of the whorls: genetic interactions controlling flower development". Nature. 353 (6339): 31–7. Bibcode:1991Natur.353...31C. doi:10.1038/353031a0. PMID 1715520. S2CID 4276098.
- ^ Tsukaya H (7 June 2013). "Leaf development". The Arabidopsis Book. 11: e0163. doi:10.1199/tab.0163. PMC 3711357. PMID 23864837.
- ^ Turner S, Sieburth LE (22 March 2003). "Vascular patterning". The Arabidopsis Book. 2: e0073. doi:10.1199/tab.0073. PMC 3243335. PMID 22303224.
- ^ Efroni I, Eshed Y, Lifschitz E (April 2010). "Morphogenesis of simple and compound leaves: a critical review". The Plant Cell. 22 (4): 1019–32. doi:10.1105/tpc.109.073601. PMC 2879760. PMID 20435903.
- ^ Moreno N, Bougourd S, Haseloff J and Fiejo JA. 2006. Chapter 44: Imaging Plant Cells. In: Pawley JB (Editor). Handbook of Biological Confocal Microscopy - 3rd edition. SpringerScience+Business Media, New York. p769-787
- ^ Shaw SL (February 2006). "Imaging the live plant cell". The Plant Journal. 45 (4): 573–98. doi:10.1111/j.1365-313X.2006.02653.x. PMID 16441350.
- ^ a b c d e f g Simpson, Gordon G.; Dean, Caroline (12 April 2002). "Arabidopsis, the Rosetta Stone of Flowering Time?". Science. 296 (5566). American Association for the Advancement of Science (AAAS): 285–289. Bibcode:2002Sci...296..285S. CiteSeerX 10.1.1.991.2232. doi:10.1126/science.296.5566.285. ISSN 0036-8075. PMID 11951029.
- ^ Friedman, Jannice (2 November 2020). "The Evolution of Annual and Perennial Plant Life Histories: Ecological Correlates and Genetic Mechanisms". Annual Review of Ecology, Evolution, and Systematics. 51 (1). Annual Reviews: 461–481. doi:10.1146/annurev-ecolsys-110218-024638. ISSN 1543-592X. S2CID 225237602.
- ^ Whittaker, Charles; Dean, Caroline (6 October 2017). "The FLC Locus: A Platform for Discoveries in Epigenetics and Adaptation". Annual Review of Cell and Developmental Biology. 33 (1). Annual Reviews: 555–575. doi:10.1146/annurev-cellbio-100616-060546. ISSN 1081-0706. PMID 28693387.
- ^ Sullivan JA, Deng XW (August 2003). "From seed to seed: the role of photoreceptors in Arabidopsis development". Developmental Biology. 260 (2): 289–97. doi:10.1016/S0012-1606(03)00212-4. PMID 12921732.
- ^ Más P (2005). "Circadian clock signaling in Arabidopsis thaliana: from gene expression to physiology and development". The International Journal of Developmental Biology. 49 (5–6): 491–500. doi:10.1387/ijdb.041968pm. PMID 16096959.
- ^ Scialdone A, Mugford ST, Feike D, Skeffington A, Borrill P, Graf A, Smith AM, Howard M (June 2013). "Arabidopsis plants perform arithmetic division to prevent starvation at night". eLife. 2: e00669. arXiv:1306.5148. doi:10.7554/eLife.00669. PMC 3691572. PMID 23805380.
- ^ Ruppel NJ, Hangarter RP, Kiss JZ (February 2001). "Red-light-induced positive phototropism in Arabidopsis roots". Planta. 212 (3): 424–30. Bibcode:2001Plant.212..424R. doi:10.1007/s004250000410. PMID 11289607. S2CID 28410755.
- ^ "Plants that Glow in the Dark" Archived 3 February 2014 at the Wayback Machine, Bioresearch Online, 18 May 2000
- ^ a b Braam, Janet; Davis, Ronald W. (9 February 1990). "Rain-, wind-, and touch-induced expression of calmodulin and calmodulin-related genes in Arabidopsis". Cell. 60 (3): 357–364. doi:10.1016/0092-8674(90)90587-5. ISSN 0092-8674. PMID 2302732. S2CID 38574940.
- ^ Pigliucci, Massimo (May 2002). "Touchy and Bushy: Phenotypic Plasticity and Integration in Response to Wind Stimulation inArabidopsis thaliana". International Journal of Plant Sciences. 163 (3): 399–408. doi:10.1086/339158. ISSN 1058-5893. S2CID 84173889.
- ^ a b Letzter, Rafi (4 January 2019). "There Are Plants and Animals on the Moon Now (Because of China)". Space.com. Archived from the original on 15 January 2019. Retrieved 15 January 2019.
- ^ Connor, Neil (13 April 2018). "China plans to grow flowers and silkworms on the dark side of the moon". The Telegraph. ISSN 0307-1235. Archived from the original on 12 January 2022. Retrieved 15 January 2019.
- ^ a b Lacchini, Elia; Goossens, Alain (2020). "Combinatorial Control of Plant Specialized Metabolism: Mechanisms, Functions, and Consequences". Annual Review of Cell and Developmental Biology. 36 (1). Annual Reviews: 291–313. doi:10.1146/annurev-cellbio-011620-031429. ISSN 1081-0706. PMID 32559387. S2CID 219947907.
- ^ Hassani, M.A., Durán, P. and Hacquard, S. (2018) "Microbial interactions within the plant holobiont". Microbiome, 6(1): 58. doi:10.1186/s40168-018-0445-0. Material was copied from this source, which is available under a Creative Commons Attribution 4.0 International License Archived 16 October 2017 at the Wayback Machine
- ^ Collins NC, Thordal-Christensen H, Lipka V, Bau S, Kombrink E, Qiu JL, Hückelhoven R, Stein M, Freialdenhoven A, Somerville SC, Schulze-Lefert P (October 2003). "SNARE-protein-mediated disease resistance at the plant cell wall". Nature. 425 (6961): 973–7. Bibcode:2003Natur.425..973C. doi:10.1038/nature02076. PMID 14586469. S2CID 4408024.
- ^ Lipka V, Dittgen J, Bednarek P, Bhat R, Wiermer M, Stein M, Landtag J, Brandt W, Rosahl S, Scheel D, Llorente F, Molina A, Parker J, Somerville S, Schulze-Lefert P (November 2005). "Pre- and postinvasion defenses both contribute to nonhost resistance in Arabidopsis". Science. 310 (5751): 1180–3. Bibcode:2005Sci...310.1180L. doi:10.1126/science.1119409. hdl:11858/00-001M-0000-0012-3A32-0. PMID 16293760. S2CID 35317665. Archived from the original on 11 March 2022. Retrieved 5 September 2019.
- ^ Stein M, Dittgen J, Sánchez-Rodríguez C, Hou BH, Molina A, Schulze-Lefert P, Lipka V, Somerville S (March 2006). "Arabidopsis PEN3/PDR8, an ATP binding cassette transporter, contributes to nonhost resistance to inappropriate pathogens that enter by direct penetration". The Plant Cell. 18 (3): 731–46. doi:10.1105/tpc.105.038372. PMC 1383646. PMID 16473969.
- ^ Knepper C, Day B (March 2010). "From perception to activation: the molecular-genetic and biochemical landscape of disease resistance signaling in plants". The Arabidopsis Book. 8: e012. doi:10.1199/tab.0124. PMC 3244959. PMID 22303251.
- ^ Gómez-Gómez L, Felix G, Boller T (May 1999). "A single locus determines sensitivity to bacterial flagellin in Arabidopsis thaliana". The Plant Journal. 18 (3): 277–84. doi:10.1046/j.1365-313X.1999.00451.x. PMID 10377993.
- ^ Gómez-Gómez L, Boller T (June 2000). "FLS2: an LRR receptor-like kinase involved in the perception of the bacterial elicitor flagellin in Arabidopsis". Molecular Cell. 5 (6): 1003–11. doi:10.1016/S1097-2765(00)80265-8. PMID 10911994.
- ^ Urban, L.; Chabane Sari, D.; Orsal, B.; Lopes, M.; Miranda, R.; Aarrouf, J. (2018). "UV-C light and pulsed light as alternatives to chemical and biological elicitors for stimulating plant natural defenses against fungal diseases". Scientia Horticulturae. 235. Elsevier: 452–459. Bibcode:2018ScHor.235..452U. doi:10.1016/j.scienta.2018.02.057. ISSN 0304-4238. S2CID 90436989.
- ^ Zipfel C, Kunze G, Chinchilla D, Caniard A, Jones JD, Boller T, Felix G (May 2006). "Perception of the bacterial PAMP EF-Tu by the receptor EFR restricts Agrobacterium-mediated transformation". Cell. 125 (4): 749–60. doi:10.1016/j.cell.2006.03.037. PMID 16713565. S2CID 6856390.
- ^ Lacombe S, Rougon-Cardoso A, Sherwood E, Peeters N, Dahlbeck D, van Esse HP, Smoker M, Rallapalli G, Thomma BP, Staskawicz B, Jones JD, Zipfel C (April 2010). "Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance". Nature Biotechnology. 28 (4): 365–9. doi:10.1038/nbt.1613. PMID 20231819. S2CID 7260214.,
- ^ Zhang J, Zhou JM (September 2010). "Plant immunity triggered by microbial molecular signatures". Molecular Plant. 3 (5): 783–93. doi:10.1093/mp/ssq035. PMID 20713980.
- ^ Kunkel BN, Bent AF, Dahlbeck D, Innes RW, Staskawicz BJ (August 1993). "RPS2, an Arabidopsis disease resistance locus specifying recognition of Pseudomonas syringae strains expressing the avirulence gene avrRpt2". The Plant Cell. 5 (8): 865–75. doi:10.1105/tpc.5.8.865. PMC 160322. PMID 8400869.
- ^ Axtell MJ, Staskawicz BJ (February 2003). "Initiation of RPS2-specified disease resistance in Arabidopsis is coupled to the AvrRpt2-directed elimination of RIN4". Cell. 112 (3): 369–77. doi:10.1016/S0092-8674(03)00036-9. PMID 12581526. S2CID 1497625.
- ^ Cao H, Bowling SA, Gordon AS, Dong X (November 1994). "Characterization of an Arabidopsis Mutant That Is Nonresponsive to Inducers of Systemic Acquired Resistance". The Plant Cell. 6 (11): 1583–1592. doi:10.1105/tpc.6.11.1583. PMC 160545. PMID 12244227.
- ^ Mou Z, Fan W, Dong X (June 2003). "Inducers of plant systemic acquired resistance regulate NPR1 function through redox changes". Cell. 113 (7): 935–44. doi:10.1016/S0092-8674(03)00429-X. PMID 12837250. S2CID 1562690.
- ^ Johnson C, Boden E, Arias J (August 2003). "Salicylic acid and NPR1 induce the recruitment of trans-activating TGA factors to a defense gene promoter in Arabidopsis". The Plant Cell. 15 (8): 1846–58. doi:10.1105/tpc.012211. PMC 167174. PMID 12897257.
- ^ a b c d e Delaney TP, Uknes S, Vernooij B, Friedrich L, Weymann K, Negrotto D, Gaffney T, Gut-Rella M, Kessmann H, Ward E, Ryals J (November 1994). "A central role of salicylic Acid in plant disease resistance". Science. 266 (5188): 1247–50. Bibcode:1994Sci...266.1247D. doi:10.1126/science.266.5188.1247. PMID 17810266. S2CID 15507678.
- ^ Ben Khaled, Sara; Postma, Jelle; Robatzek, Silke (4 August 2015). "A Moving View: Subcellular Trafficking Processes in Pattern Recognition Receptor–Triggered Plant Immunity". Annual Review of Phytopathology. 53 (1). Annual Reviews: 379–402. doi:10.1146/annurev-phyto-080614-120347. ISSN 0066-4286. PMID 26243727.
- ^ Bent AF, Kunkel BN, Dahlbeck D, Brown KL, Schmidt R, Giraudat J, Leung J, Staskawicz BJ (September 1994). "RPS2 of Arabidopsis thaliana: a leucine-rich repeat class of plant disease resistance genes". Science. 265 (5180): 1856–60. Bibcode:1994Sci...265.1856B. doi:10.1126/science.8091210. PMID 8091210.
- ^ a b c d Zipfel C, Robatzek S, Navarro L, Oakeley EJ, Jones JD, Felix G, Boller T (April 2004). "Bacterial disease resistance in Arabidopsis through flagellin perception". Nature. 428 (6984): 764–7. Bibcode:2004Natur.428..764Z. doi:10.1038/nature02485. PMID 15085136. S2CID 4332562.
- ^ a b Lawton K, Friedrich L, Hunt M (1996). "Benzothiadizaole induces disease resistance by a citation of the systemic acquired resistance signal transduction pathway". The Plant Journal. 10 (1): 71–82. doi:10.1046/j.1365-313x.1996.10010071.x. PMID 8758979.
- ^ Fridman, Eyal (2015). "Consequences of hybridization and heterozygosity on plant vigor and phenotypic stability". Plant Science. 232. Elsevier: 35–40. Bibcode:2015PlnSc.232...35F. doi:10.1016/j.plantsci.2014.11.014. ISSN 0168-9452. PMID 25617321.
- ^ Lapin, Dmitry; Bhandari, Deepak D.; Parker, Jane E. (25 August 2020). "Origins and Immunity Networking Functions of EDS1 Family Proteins". Annual Review of Phytopathology. 58 (1). Annual Reviews: 253–276. doi:10.1146/annurev-phyto-010820-012840. hdl:1874/413668. ISSN 0066-4286. PMID 32396762. S2CID 218617308.
- ^ Pottinger, Sarah E.; Innes, Roger W. (25 August 2020). "RPS5-Mediated Disease Resistance: Fundamental Insights and Translational Applications". Annual Review of Phytopathology. 58 (1). Annual Reviews: 139–160. doi:10.1146/annurev-phyto-010820-012733. ISSN 0066-4286. PMID 32284014. S2CID 215757180.
- ^ Link BM, Busse JS, Stankovic B (2014). "Seed-to-Seed-to-Seed Growth and Development of Arabidopsis in Microgravity". Astrobiology. 14 (10): 866–875. Bibcode:2014AsBio..14..866L. doi:10.1089/ast.2014.1184. PMC 4201294. PMID 25317938.
- ^ Ferl RJ, Paul AL (April 2010). "Lunar plant biology--a review of the Apollo era". Astrobiology. 10 (3): 261–74. Bibcode:2010AsBio..10..261F. doi:10.1089/ast.2009.0417. PMID 20446867.
- ^ Yetisen AK, Jiang L, Cooper JR, Qin Y, Palanivelu R, Zohar Y (May 2011). "A microsystem-based assay for studying pollen tube guidance in plant reproduction". J. Micromech. Microeng. 25 (5): 054018. Bibcode:2011JMiMi..21e4018Y. doi:10.1088/0960-1317/21/5/054018. S2CID 12989263.
- ^ "NASA-funded study breaks new ground in plant research". NASA. 12 May 2022. Archived from the original on 12 May 2022. Retrieved 13 May 2022.
- ^ Abbott RJ, Gomes MF (1989). "Population genetic structure and outcrossing rate of Arabidopsis thaliana (L.) Heynh". Heredity. 62 (3): 411–418. doi:10.1038/hdy.1989.56.
- ^ Tang C, Toomajian C, Sherman-Broyles S, Plagnol V, Guo YL, Hu TT, Clark RM, Nasrallah JB, Weigel D, Nordborg M (August 2007). "The evolution of selfing in Arabidopsis thaliana". Science. 317 (5841): 1070–2. Bibcode:2007Sci...317.1070T. doi:10.1126/science.1143153. PMID 17656687. S2CID 45853624.
- ^ Bernstein H; Byerly HC; Hopf FA; Michod RE (1985). "Genetic damage, mutation, and the evolution of sex". Science. 229 (4719): 1277–81. Bibcode:1985Sci...229.1277B. doi:10.1126/science.3898363. PMID 3898363
- ^ "TAIR - Gene Expression - Microarray - Public Datasets". Archived from the original on 4 December 2021. Retrieved 4 December 2021.
- ^ "Genome research inspires new bridge at Harlow Carr". The Garden (September 2021): 97. 2021.
External links
edit- Arabidopsis transcriptional regulatory map
- The Arabidopsis Information Resource (TAIR)
- Salk Institute Genomic Analysis Laboratory Archived 8 March 2021 at the Wayback Machine
- What Makes Plants Grow? The Arabidopsis genome knows Featured article in Genome News Network
- The Arabidopsis book - A comprehensive review published yearly related to research in Arabidopsis
- A. thaliana protein abundance
- The Arabidopsis Information Portal (Araport)