In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of itself, such as tRNA or rRNA. Promoters are located near the transcription start sites of genes, upstream on the DNA (towards the 5' region of the sense strand). Promoters can be about 100–1000 base pairs long, the sequence of which is highly dependent on the gene and product of transcription, type or class of RNA polymerase recruited to the site, and species of organism.[1][2]
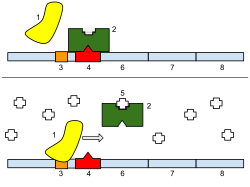
Bottom: The gene is turned on. Lactose is inhibiting the repressor, allowing the RNA polymerase to bind with the promoter and express the genes, which synthesize lactase. Eventually, the lactase will digest all of the lactose, until there is none to bind to the repressor. The repressor will then bind to the operator, stopping the manufacture of lactase.
Promoters control gene expression in bacteria and eukaryotes.[3] RNA polymerase must attach to DNA near a gene for transcription to occur. Promoter DNA sequences provide an enzyme binding site. The -10 sequence is TATAAT. -35 sequences are conserved on average, but not in most promoters.
Artificial promoters with conserved -10 and -35 elements transcribe more slowly. All DNAs have "Closely spaced promoters". Divergent, tandem, and convergent orientations are possible. Two closely spaced promoters will likely interfere. Regulatory elements can be several kilobases away from the transcriptional start site in gene promoters (enhancers).
In eukaryotes, the transcriptional complex can bend DNA, allowing regulatory sequences to be placed far from the transcription site. The distal promoter is upstream of the gene and may contain additional regulatory elements with a weaker influence. RNA polymerase II (RNAP II) bound to the transcription start site promoter can start mRNA synthesis. It also typically contains CpG islands, a TATA box, and TFIIB recognition elements.
Hypermethylation downregulates both genes, while demethylation upregulates them. Non-coding RNAs are linked to mRNA promoter regions. Subgenomic promoters range from 24 to 100 nucleotides (Beet necrotic yellow vein virus). Gene expression depends on promoter binding. Unwanted gene changes can increase a cell's cancer risk.
MicroRNA promoters often contain CpG islands. DNA methylation forms 5-methylcytosines at the 5' pyrimidine ring of CpG cytosine residues. Some cancer genes are silenced by mutation, but most are silenced by DNA methylation. Others are regulated promoters. Selection may favor less energetic transcriptional binding.
Variations in promoters or transcription factors cause some diseases. Misunderstandings can result from using a canonical sequence to describe a promoter.
Overview
editFor transcription to take place, the enzyme that synthesizes RNA, known as RNA polymerase, must attach to the DNA near a gene. Promoters contain specific DNA sequences such as response elements that provide a secure initial binding site for RNA polymerase and for proteins called transcription factors that recruit RNA polymerase. These transcription factors have specific activator or repressor sequences of corresponding nucleotides that attach to specific promoters and regulate gene expression.[citation needed]
- In bacteria
- The promoter is recognized by RNA polymerase and an associated sigma factor, which in turn are often brought to the promoter DNA by an activator protein's binding to its own DNA binding site nearby.
- In eukaryotes
- The process is more complicated, and at least seven different factors are necessary for the binding of an RNA polymerase II to the promoter.
Promoters represent critical elements that can work in concert with other regulatory regions (enhancers, silencers, boundary elements/insulators) to direct the level of transcription of a given gene. A promoter is induced in response to changes in abundance or conformation of regulatory proteins in a cell, which enable activating transcription factors to recruit RNA polymerase.[4][5]
Given the short sequences of most promoter elements, promoters can rapidly evolve from random sequences. For instance, in E. coli, ~60% of random sequences can evolve expression levels comparable to the wild-type lac promoter with only one mutation, and that ~10% of random sequences can serve as active promoters even without evolution.[6]
Identification of relative location
editAs promoters are typically immediately adjacent to the gene in question, positions in the promoter are designated relative to the transcriptional start site, where transcription of DNA begins for a particular gene (i.e., positions upstream are negative numbers counting back from -1, for example -100 is a position 100 base pairs upstream).[citation needed]
Elements
edit
Bacterial
editIn bacteria, the promoter contains two short sequence elements approximately 10 (Pribnow Box) and 35 nucleotides upstream from the transcription start site.[2]
- The sequence at -10 (the -10 element) has the consensus sequence TATAAT.
- The sequence at -35 (the -35 element) has the consensus sequence TTGACA.
- The above consensus sequences, while conserved on average, are not found intact in most promoters. On average, only 3 to 4 of the 6 base pairs in each consensus sequence are found in any given promoter. Few natural promoters have been identified to date that possess intact consensus sequences at both the -10 and -35; artificial promoters with complete conservation of the -10 and -35 elements have been found to transcribe at lower frequencies than those with a few mismatches with the consensus.
- The optimal spacing between the -35 and -10 sequences is 17 bp.
- Some promoters contain one or more upstream promoter element (UP element) subsites[7] (consensus sequence 5'-AAAAAARNR-3' when centered in the -42 region; consensus sequence 5'-AWWWWWTTTTT-3' when centered in the -52 region; W = A or T; R = A or G; N = any base).[8]
The above promoter sequences are recognized only by RNA polymerase holoenzyme containing sigma-70. RNA polymerase holoenzymes containing other sigma factors recognize different core promoter sequences.
← upstream downstream →
5'-XXXXXXXPPPPPPXXXXXXPPPPPPXXXXGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGGXXXX-3'
-35 -10 Gene to be transcribed
Probability of occurrence of each nucleotide
editfor -10 sequence T A T A A T 77% 76% 60% 61% 56% 82%
for -35 sequence T T G A C A 69% 79% 61% 56% 54% 54%
Bidirectional (prokaryotic)
editPromoters can be very closely located in the DNA. Such "closely spaced promoters" have been observed in the DNAs of all life forms, from humans[9] to prokaryotes[10] and are highly conserved.[11] Therefore, they may provide some (presently unknown) advantages. These pairs of promoters can be positioned in divergent, tandem, and convergent directions. They can also be regulated by transcription factors and differ in various features, such as the nucleotide distance between them, the two promoter strengths, etc. The most important aspect of two closely spaced promoters is that they will, most likely, interfere with each other. Several studies have explored this using both analytical and stochastic models.[12][13][14] There are also studies that measured gene expression in synthetic genes or from one to a few genes controlled by bidirectional promoters.[15]
More recently, one study measured most genes controlled by tandem promoters in E. coli.[16] In that study, two main forms of interference were measured. One is when an RNAP is on the downstream promoter, blocking the movement of RNAPs elongating from the upstream promoter. The other is when the two promoters are so close that when an RNAP sits on one of the promoters, it blocks any other RNAP from reaching the other promoter. These events are possible because the RNAP occupies several nucleotides when bound to the DNA, including in transcription start sites. Similar events occur when the promoters are in divergent and convergent formations. The possible events also depend on the distance between them.
Eukaryotic
editGene promoters are typically located upstream of the gene and can have regulatory elements several kilobases away from the transcriptional start site (enhancers). In eukaryotes, the transcriptional complex can cause the DNA to bend back on itself, which allows for placement of regulatory sequences far from the actual site of transcription. Eukaryotic RNA-polymerase-II-dependent promoters can contain a TATA box (consensus sequence TATAAA), which is recognized by the general transcription factor TATA-binding protein (TBP); and a B recognition element (BRE), which is recognized by the general transcription factor TFIIB.[17][18][19] The TATA element and BRE typically are located close to the transcriptional start site (typically within 30 to 40 base pairs).
Eukaryotic promoter regulatory sequences typically bind proteins called transcription factors that are involved in the formation of the transcriptional complex. An example is the E-box (sequence CACGTG), which binds transcription factors in the basic helix-loop-helix (bHLH) family (e.g. BMAL1-Clock, cMyc).[20] Some promoters that are targeted by multiple transcription factors might achieve a hyperactive state, leading to increased transcriptional activity.[21]
- Core promoter – the minimal portion of the promoter required to properly initiate transcription[17]
- Includes the transcription start site (TSS) and elements directly upstream
- A binding site for RNA polymerase
- RNA polymerase I: transcribes genes encoding 18S, 5.8S and 28S ribosomal RNAs
- RNA polymerase II: transcribes genes encoding messenger RNA and certain small nuclear RNAs and microRNA
- RNA polymerase III: transcribes genes encoding transfer RNA, 5s ribosomal RNAs and other small RNAs
- General transcription factor binding sites, e.g. TATA box, B recognition element.
- Many other elements/motifs may be present. There is no such thing as a set of "universal elements" found in every core promoter.[22]
- Proximal promoter – the proximal sequence upstream of the gene that tends to contain primary regulatory elements
- Approximately 250 base pairs upstream of the start site
- Specific transcription factor binding sites
- Distal promoter – the distal sequence upstream of the gene that may contain additional regulatory elements, often with a weaker influence than the proximal promoter
- Anything further upstream (but not an enhancer or other regulatory region whose influence is positional/orientation independent)
- Specific transcription factor binding sites
Mammalian promoters
editThis section may require cleanup to meet Wikipedia's quality standards. The specific problem is: Text about mammals highly duplicated among uses of the same picture -- can we make a "canonical" version and redirect people there? (September 2021) |
Up-regulated expression of genes in mammals is initiated when signals are transmitted to the promoters associated with the genes. Promoter DNA sequences may include different elements such as CpG islands (present in about 70% of promoters), a TATA box (present in about 24% of promoters), initiator (Inr) (present in about 49% of promoters), upstream and downstream TFIIB recognition elements (BREu and BREd) (present in about 22% of promoters), and downstream core promoter element (DPE) (present in about 12% of promoters).[23] The presence of multiple methylated CpG sites in CpG islands of promoters causes stable silencing of genes.[24] However, the presence or absence of the other elements have relatively small effects on gene expression in experiments.[25] Two sequences, the TATA box and Inr, caused small but significant increases in expression (45% and 28% increases, respectively). The BREu and the BREd elements significantly decreased expression by 35% and 20%, respectively, and the DPE element had no detected effect on expression.[25]
Cis-regulatory modules that are localized in DNA regions distant from the promoters of genes can have very large effects on gene expression, with some genes undergoing up to 100-fold increased expression due to such a cis-regulatory module.[26] These cis-regulatory modules include enhancers, silencers, insulators and tethering elements.[27] Among this constellation of elements, enhancers and their associated transcription factors have a leading role in the regulation of gene expression.[28]
Enhancers are regions of the genome that are major gene-regulatory elements. Enhancers control cell-type-specific gene expression programs, most often by looping through long distances to come in physical proximity with the promoters of their target genes.[29] In a study of brain cortical neurons, 24,937 loops were found, bringing enhancers to promoters.[26] Multiple enhancers, each often at tens or hundred of thousands of nucleotides distant from their target genes, loop to their target gene promoters and coordinate with each other to control expression of their common target gene.[29]
The schematic illustration in this section shows an enhancer looping around to come into close physical proximity with the promoter of a target gene. The loop is stabilized by a dimer of a connector protein (e.g. dimer of CTCF or YY1), with one member of the dimer anchored to its binding motif on the enhancer and the other member anchored to its binding motif on the promoter (represented by the red zigzags in the illustration).[30] Several cell function specific transcription factors (there are about 1,600 transcription factors in a human cell[31]) generally bind to specific motifs on an enhancer[32] and a small combination of these enhancer-bound transcription factors, when brought close to a promoter by a DNA loop, govern the level of transcription of the target gene. Mediator (coactivator) (a complex usually consisting of about 26 proteins in an interacting structure) communicates regulatory signals from enhancer DNA-bound transcription factors directly to the RNA polymerase II (pol II) enzyme bound to the promoter.[33]
Enhancers, when active, are generally transcribed from both strands of DNA with RNA polymerases acting in two different directions, producing two eRNAs as illustrated in the Figure.[34] An inactive enhancer may be bound by an inactive transcription factor. Phosphorylation of the transcription factor may activate it and that activated transcription factor may then activate the enhancer to which it is bound (see small red star representing phosphorylation of transcription factor bound to enhancer in the illustration).[35] An activated enhancer begins transcription of its RNA before activating a promoter to initiate transcription of messenger RNA from its target gene.[36]
Bidirectional (mammalian)
editBidirectional promoters are short (<1 kbp) intergenic regions of DNA between the 5' ends of the genes in a bidirectional gene pair.[37] A "bidirectional gene pair" refers to two adjacent genes coded on opposite strands, with their 5' ends oriented toward one another.[38] The two genes are often functionally related, and modification of their shared promoter region allows them to be co-regulated and thus co-expressed.[39] Bidirectional promoters are a common feature of mammalian genomes.[40] About 11% of human genes are bidirectionally paired.[37]
Bidirectionally paired genes in the Gene Ontology database shared at least one database-assigned functional category with their partners 47% of the time.[41] Microarray analysis has shown bidirectionally paired genes to be co-expressed to a higher degree than random genes or neighboring unidirectional genes.[37] Although co-expression does not necessarily indicate co-regulation, methylation of bidirectional promoter regions has been shown to downregulate both genes, and demethylation to upregulate both genes.[42] There are exceptions to this, however. In some cases (about 11%), only one gene of a bidirectional pair is expressed.[37] In these cases, the promoter is implicated in suppression of the non-expressed gene. The mechanism behind this could be competition for the same polymerases, or chromatin modification. Divergent transcription could shift nucleosomes to upregulate transcription of one gene, or remove bound transcription factors to downregulate transcription of one gene.[43]
Some functional classes of genes are more likely to be bidirectionally paired than others. Genes implicated in DNA repair are five times more likely to be regulated by bidirectional promoters than by unidirectional promoters. Chaperone proteins are three times more likely, and mitochondrial genes are more than twice as likely. Many basic housekeeping and cellular metabolic genes are regulated by bidirectional promoters.[37] The overrepresentation of bidirectionally paired DNA repair genes associates these promoters with cancer. Forty-five percent of human somatic oncogenes seem to be regulated by bidirectional promoters – significantly more than non-cancer causing genes. Hypermethylation of the promoters between gene pairs WNT9A/CD558500, CTDSPL/BC040563, and KCNK15/BF195580 has been associated with tumors.[42]
Certain sequence characteristics have been observed in bidirectional promoters, including a lack of TATA boxes, an abundance of CpG islands, and a symmetry around the midpoint of dominant Cs and As on one side and Gs and Ts on the other. A motif with the consensus sequence of TCTCGCGAGA, also called the CGCG element, was recently shown to drive PolII-driven bidirectional transcription in CpG islands.[44] CCAAT boxes are common, as they are in many promoters that lack TATA boxes. In addition, the motifs NRF-1, GABPA, YY1, and ACTACAnnTCCC are represented in bidirectional promoters at significantly higher rates than in unidirectional promoters. The absence of TATA boxes in bidirectional promoters suggests that TATA boxes play a role in determining the directionality of promoters, but counterexamples of bidirectional promoters do possess TATA boxes and unidirectional promoters without them indicates that they cannot be the only factor.[45]
Although the term "bidirectional promoter" refers specifically to promoter regions of mRNA-encoding genes, luciferase assays have shown that over half of human genes do not have a strong directional bias. Research suggests that non-coding RNAs are frequently associated with the promoter regions of mRNA-encoding genes. It has been hypothesized that the recruitment and initiation of RNA polymerase II usually begins bidirectionally, but divergent transcription is halted at a checkpoint later during elongation. Possible mechanisms behind this regulation include sequences in the promoter region, chromatin modification, and the spatial orientation of the DNA.[43]
Subgenomic
editA subgenomic promoter is a promoter added to a virus for a specific heterologous gene, resulting in the formation of mRNA for that gene alone. Many positive-sense RNA viruses produce these subgenomic mRNAs (sgRNA) as one of the common infection techniques used by these viruses and generally transcribe late viral genes. Subgenomic promoters range from 24 nucleotide (Sindbis virus) to over 100 nucleotides (Beet necrotic yellow vein virus) and are usually found upstream of the transcription start.[46]
Detection
editA wide variety of algorithms have been developed to facilitate detection of promoters in genomic sequence, and promoter prediction is a common element of many gene prediction methods. A promoter region is located before the -35 and -10 Consensus sequences. The closer the promoter region is to the consensus sequences the more often transcription of that gene will take place. There is not a set pattern for promoter regions as there are for consensus sequences.
Binding
editThe initiation of the transcription is a multistep sequential process that involves several mechanisms: promoter location, initial reversible binding of RNA polymerase, conformational changes in RNA polymerase, conformational changes in DNA, binding of nucleoside triphosphate (NTP) to the functional RNA polymerase-promoter complex, and nonproductive and productive initiation of RNA synthesis.[47][2]
The promoter binding process is crucial in the understanding of the process of gene expression. Tuning synthetic genetic systems relies on precisely engineered synthetic promoters with known levels of transcription rates.[2]
Location
editAlthough RNA polymerase holoenzyme shows high affinity to non-specific sites of the DNA, this characteristic does not allow us to clarify the process of promoter location.[48] This process of promoter location has been attributed to the structure of the holoenzyme to DNA and sigma 4 to DNA complexes.[49]
Diseases associated with aberrant function
editMost diseases are heterogeneous in cause, meaning that one "disease" is often many different diseases at the molecular level, though symptoms exhibited and response to treatment may be identical. How diseases of different molecular origin respond to treatments is partially addressed in the discipline of pharmacogenomics.
Not listed here are the many kinds of cancers involving aberrant transcriptional regulation owing to creation of chimeric genes through pathological chromosomal translocation. Importantly, intervention in the number or structure of promoter-bound proteins is one key to treating a disease without affecting expression of unrelated genes sharing elements with the target gene.[50] Some genes whose change is not desirable are capable of influencing the potential of a cell to become cancerous.[51]
CpG islands in promoters
editIn humans, about 70% of promoters located near the transcription start site of a gene (proximal promoters) contain a CpG island.[52][53] CpG islands are generally 200 to 2000 base pairs long, have a C:G base pair content >50%, and have regions of DNA where a cytosine nucleotide is followed by a guanine nucleotide and this occurs frequently in the linear sequence of bases along its 5' → 3' direction.
Distal promoters also frequently contain CpG islands, such as the promoter of the DNA repair gene ERCC1, where the CpG island-containing promoter is located about 5,400 nucleotides upstream of the coding region of the ERCC1 gene.[54] CpG islands also occur frequently in promoters for functional noncoding RNAs such as microRNAs.
Methylation of CpG islands stably silences genes
editIn humans, DNA methylation occurs at the 5' position of the pyrimidine ring of the cytosine residues within CpG sites to form 5-methylcytosines. The presence of multiple methylated CpG sites in CpG islands of promoters causes stable silencing of genes.[24] Silencing of a gene may be initiated by other mechanisms, but this is often followed by methylation of CpG sites in the promoter CpG island to cause the stable silencing of the gene.[24]
Promoter CpG hyper/hypo-methylation in cancer
editGenerally, in progression to cancer, hundreds of genes are silenced or activated. Although silencing of some genes in cancers occurs by mutation, a large proportion of carcinogenic gene silencing is a result of altered DNA methylation (see DNA methylation in cancer). DNA methylation causing silencing in cancer typically occurs at multiple CpG sites in the CpG islands that are present in the promoters of protein coding genes.
Altered expressions of microRNAs also silence or activate many genes in progression to cancer (see microRNAs in cancer). Altered microRNA expression occurs through hyper/hypo-methylation of CpG sites in CpG islands in promoters controlling transcription of the microRNAs.
Silencing of DNA repair genes through methylation of CpG islands in their promoters appears to be especially important in progression to cancer (see methylation of DNA repair genes in cancer).
Canonical sequences and wild-type
editThe usage of the term canonical sequence to refer to a promoter is often problematic, and can lead to misunderstandings about promoter sequences. Canonical implies perfect, in some sense.
In the case of a transcription factor binding site, there may be a single sequence that binds the protein most strongly under specified cellular conditions. This might be called canonical.
However, natural selection may favor less energetic binding as a way of regulating transcriptional output. In this case, we may call the most common sequence in a population the wild-type sequence. It may not even be the most advantageous sequence to have under prevailing conditions.
Recent evidence also indicates that several genes (including the proto-oncogene c-myc) have G-quadruplex motifs as potential regulatory signals.
Synthetic promoter design and engineering
editPromoters are important gene regulatory elements used in tuning synthetically designed genetic circuits and metabolic networks. For example, to overexpress an important gene in a network, to yield higher production of target protein, synthetic biologists design promoters to upregulate its expression. Automated algorithms can be used to design neutral DNA or insulators that do not trigger gene expression of downstream sequences.[55][2]
Diseases that may be associated with variations
editSome cases of many genetic diseases are associated with variations in promoters or transcription factors.
Examples include:
Constitutive vs regulated
editSome promoters are called constitutive as they are active in all circumstances in the cell, while others are regulated, becoming active in the cell only in response to specific stimuli.
Tissue-specific promoter
editA tissue-specific promoter is a promoter that has activity in only certain cell types.
Use of the term
editWhen referring to a promoter some authors actually mean promoter + operator; i.e., the lac promoter is IPTG inducible, meaning that besides the lac promoter, the lac operon is also present. If the lac operator were not present the IPTG would not have an inducible effect.[citation needed] Another example is the Tac-Promoter system (Ptac). Notice how tac is written as a tac promoter, while in fact tac is actually both a promoter and an operator.[60]
See also
editReferences
edit- ^ Sharan R (4 January 2007). "Analysis of Biological Networks: Transcriptional Networks – Promoter Sequence Analysis" (PDF). Tel Aviv University. Retrieved 30 December 2012.
- ^ a b c d e LaFleur TL, Hossain A, Salis HM (September 2022). "Automated model-predictive design of synthetic promoters to control transcriptional profiles in bacteria". Nature Communications. 13 (1): 5159. Bibcode:2022NatCo..13.5159L. doi:10.1038/s41467-022-32829-5. PMC 9440211. PMID 36056029.
- ^ Vaishnav ED, de Boer CG, Molinet J, Yassour M, Fan L, Adiconis X, et al. (March 2022). "The evolution, evolvability and engineering of gene regulatory DNA". Nature. 603 (7901): 455–463. Bibcode:2022Natur.603..455V. doi:10.1038/s41586-022-04506-6. PMC 8934302. PMID 35264797.
- ^ Yaniv M (September 2014). "Chromatin remodeling: from transcription to cancer". Cancer Genetics. 207 (9): 352–7. doi:10.1016/j.cancergen.2014.03.006. PMID 24825771.
- ^ Civas A, Génin P, Morin P, Lin R, Hiscott J (February 2006). "Promoter organization of the interferon-A genes differentially affects virus-induced expression and responsiveness to TBK1 and IKKepsilon". The Journal of Biological Chemistry. 281 (8): 4856–66. doi:10.1074/jbc.M506812200. PMID 16380379.
- ^ Yona AH, Alm EJ, Gore J (April 2018). "Random sequences rapidly evolve into de novo promoters". Nature Communications. 9 (1): 1530. Bibcode:2018NatCo...9.1530Y. doi:10.1038/s41467-018-04026-w. PMC 5906472. PMID 29670097.
- ^ Ross W, Gosink KK, Salomon J, Igarashi K, Zou C, Ishihama A, et al. (November 1993). "A third recognition element in bacterial promoters: DNA binding by the alpha subunit of RNA polymerase". Science. 262 (5138): 1407–1413. Bibcode:1993Sci...262.1407R. doi:10.1126/science.8248780. PMID 8248780.
- ^ Estrem ST, Ross W, Gaal T, Chen ZW, Niu W, Ebright RH, Gourse RL (August 1999). "Bacterial promoter architecture: subsite structure of UP elements and interactions with the carboxy-terminal domain of the RNA polymerase alpha subunit". Genes & Development. 13 (16): 2134–2147. doi:10.1101/gad.13.16.2134. PMC 316962. PMID 10465790.
- ^ Adachi N, Lieber MR (June 2002). "Bidirectional gene organization: a common architectural feature of the human genome". Cell. 109 (7): 807–809. doi:10.1016/S0092-8674(02)00758-4. PMID 12110178.
- ^ Herbert M, Kolb A, Buc H (May 1986). "Overlapping promoters and their control in Escherichia coli: the gal case". Proceedings of the National Academy of Sciences of the United States of America. 83 (9): 2807–2811. Bibcode:1986PNAS...83.2807H. doi:10.1073/pnas.83.9.2807. PMC 323395. PMID 3010319.
- ^ Korbel JO, Jensen LJ, von Mering C, Bork P (July 2004). "Analysis of genomic context: prediction of functional associations from conserved bidirectionally transcribed gene pairs". Nature Biotechnology. 22 (7): 911–917. doi:10.1038/nbt988. PMID 15229555. S2CID 3546895.
- ^ Sneppen K, Dodd IB, Shearwin KE, Palmer AC, Schubert RA, Callen BP, Egan JB (February 2005). "A mathematical model for transcriptional interference by RNA polymerase traffic in Escherichia coli". Journal of Molecular Biology. 346 (2): 399–409. doi:10.1016/j.jmb.2004.11.075. PMID 15670592.
- ^ Martins L, Mäkelä J, Häkkinen A, Kandhavelu M, Yli-Harja O, Fonseca JM, Ribeiro AS (May 2012). "Dynamics of transcription of closely spaced promoters in Escherichia coli, one event at a time". Journal of Theoretical Biology. 301: 83–94. Bibcode:2012JThBi.301...83M. doi:10.1016/j.jtbi.2012.02.015. PMID 22370562.
- ^ Häkkinen A, Oliveira SM, Neeli-Venkata R, Ribeiro AS (December 2019). "Transcription closed and open complex formation coordinate expression of genes with a shared promoter region". Journal of the Royal Society, Interface. 16 (161): 20190507. doi:10.1098/rsif.2019.0507. PMC 6936044. PMID 31822223.
- ^ Bordoy AE, Varanasi US, Courtney CM, Chatterjee A (December 2016). "Transcriptional Interference in Convergent Promoters as a Means for Tunable Gene Expression". ACS Synthetic Biology. 5 (12): 1331–1341. doi:10.1021/acssynbio.5b00223. PMID 27346626.
- ^ Chauhan V, Bahrudeen MN, Palma CS, Baptista IS, Almeida BL, Dash S, et al. (January 2022). "Analytical kinetic model of native tandem promoters in E. coli". PLOS Computational Biology. 18 (1): e1009824. Bibcode:2022PLSCB..18E9824C. doi:10.1371/journal.pcbi.1009824. PMC 8830795. PMID 35100257.
- ^ a b Smale ST, Kadonaga JT (2003). "The RNA polymerase II core promoter". Annual Review of Biochemistry. 72: 449–479. doi:10.1146/annurev.biochem.72.121801.161520. PMID 12651739.
- ^ Gershenzon NI, Ioshikhes IP (April 2005). "Synergy of human Pol II core promoter elements revealed by statistical sequence analysis". Bioinformatics. 21 (8): 1295–1300. doi:10.1093/bioinformatics/bti172. PMID 15572469.
- ^ Lagrange T, Kapanidis AN, Tang H, Reinberg D, Ebright RH (January 1998). "New core promoter element in RNA polymerase II-dependent transcription: sequence-specific DNA binding by transcription factor IIB". Genes & Development. 12 (1): 34–44. doi:10.1101/gad.12.1.34. PMC 316406. PMID 9420329.
- ^ Levine M, Tjian R (July 2003). "Transcription regulation and animal diversity". Nature. 424 (6945): 147–151. Bibcode:2003Natur.424..147L. doi:10.1038/nature01763. PMID 12853946. S2CID 4373712.
- ^ Liefke R, Windhof-Jaidhauser IM, Gaedcke J, Salinas-Riester G, Wu F, Ghadimi M, Dango S (June 2015). "The oxidative demethylase ALKBH3 marks hyperactive gene promoters in human cancer cells". Genome Medicine. 7 (1): 66. doi:10.1186/s13073-015-0180-0. PMC 4517488. PMID 26221185.
- ^ Juven-Gershon T, Kadonaga JT (March 2010). "Regulation of gene expression via the core promoter and the basal transcriptional machinery". Developmental Biology. 339 (2): 225–229. doi:10.1016/j.ydbio.2009.08.009. PMC 2830304. PMID 19682982.
- ^ Yang C, Bolotin E, Jiang T, Sladek FM, Martinez E (March 2007). "Prevalence of the initiator over the TATA box in human and yeast genes and identification of DNA motifs enriched in human TATA-less core promoters". Gene. 389 (1): 52–65. doi:10.1016/j.gene.2006.09.029. PMC 1955227. PMID 17123746.
- ^ a b c Bird A (January 2002). "DNA methylation patterns and epigenetic memory". Genes & Development. 16 (1): 6–21. doi:10.1101/gad.947102. PMID 11782440.
- ^ a b Weingarten-Gabbay S, Nir R, Lubliner S, Sharon E, Kalma Y, Weinberger A, Segal E (February 2019). "Systematic interrogation of human promoters". Genome Research. 29 (2): 171–183. doi:10.1101/gr.236075.118. PMC 6360817. PMID 30622120.
- ^ a b Beagan JA, Pastuzyn ED, Fernandez LR, Guo MH, Feng K, Titus KR, et al. (June 2020). "Three-dimensional genome restructuring across timescales of activity-induced neuronal gene expression". Nature Neuroscience. 23 (6): 707–717. doi:10.1038/s41593-020-0634-6. PMC 7558717. PMID 32451484.
- ^ Verheul TC, van Hijfte L, Perenthaler E, Barakat TS (2020). "The Why of YY1: Mechanisms of Transcriptional Regulation by Yin Yang 1". Frontiers in Cell and Developmental Biology. 8: 592164. doi:10.3389/fcell.2020.592164. PMC 7554316. PMID 33102493.
- ^ Spitz F, Furlong EE (September 2012). "Transcription factors: from enhancer binding to developmental control". Nature Reviews. Genetics. 13 (9): 613–626. doi:10.1038/nrg3207. PMID 22868264. S2CID 205485256.
- ^ a b Schoenfelder S, Fraser P (August 2019). "Long-range enhancer-promoter contacts in gene expression control". Nature Reviews. Genetics. 20 (8): 437–455. doi:10.1038/s41576-019-0128-0. PMID 31086298. S2CID 152283312.
- ^ Weintraub AS, Li CH, Zamudio AV, Sigova AA, Hannett NM, Day DS, et al. (December 2017). "YY1 Is a Structural Regulator of Enhancer-Promoter Loops". Cell. 171 (7): 1573–1588.e28. doi:10.1016/j.cell.2017.11.008. PMC 5785279. PMID 29224777.
- ^ Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, et al. (February 2018). "The Human Transcription Factors". Cell. 172 (4): 650–665. doi:10.1016/j.cell.2018.01.029. PMID 29425488.
- ^ Grossman SR, Engreitz J, Ray JP, Nguyen TH, Hacohen N, Lander ES (July 2018). "Positional specificity of different transcription factor classes within enhancers". Proceedings of the National Academy of Sciences of the United States of America. 115 (30): E7222–E7230. Bibcode:2018PNAS..115E7222G. doi:10.1073/pnas.1804663115. PMC 6065035. PMID 29987030.
- ^ Allen BL, Taatjes DJ (March 2015). "The Mediator complex: a central integrator of transcription". Nature Reviews. Molecular Cell Biology. 16 (3): 155–166. doi:10.1038/nrm3951. PMC 4963239. PMID 25693131.
- ^ Mikhaylichenko O, Bondarenko V, Harnett D, Schor IE, Males M, Viales RR, Furlong EE (January 2018). "The degree of enhancer or promoter activity is reflected by the levels and directionality of eRNA transcription". Genes & Development. 32 (1): 42–57. doi:10.1101/gad.308619.117. PMC 5828394. PMID 29378788.
- ^ Li QJ, Yang SH, Maeda Y, Sladek FM, Sharrocks AD, Martins-Green M (January 2003). "MAP kinase phosphorylation-dependent activation of Elk-1 leads to activation of the co-activator p300". The EMBO Journal. 22 (2): 281–291. doi:10.1093/emboj/cdg028. PMC 140103. PMID 12514134.
- ^ Carullo NV, Phillips Iii RA, Simon RC, Soto SA, Hinds JE, Salisbury AJ, et al. (September 2020). "Enhancer RNAs predict enhancer-gene regulatory links and are critical for enhancer function in neuronal systems". Nucleic Acids Research. 48 (17): 9550–9570. doi:10.1093/nar/gkaa671. PMC 7515708. PMID 32810208.
- ^ a b c d e Trinklein ND, Aldred SF, Hartman SJ, Schroeder DI, Otillar RP, Myers RM (January 2004). "An abundance of bidirectional promoters in the human genome". Genome Research. 14 (1): 62–66. doi:10.1101/gr.1982804. PMC 314279. PMID 14707170.
- ^ Yang MQ, Koehly LM, Elnitski LL (April 2007). "Comprehensive annotation of bidirectional promoters identifies co-regulation among breast and ovarian cancer genes". PLOS Computational Biology. 3 (4): e72. Bibcode:2007PLSCB...3...72Y. doi:10.1371/journal.pcbi.0030072. PMC 1853124. PMID 17447839.
- ^ Adachi N, Lieber MR (June 2002). "Bidirectional gene organization: a common architectural feature of the human genome". Cell. 109 (7): 807–809. doi:10.1016/S0092-8674(02)00758-4. PMID 12110178. S2CID 8556921.
- ^ Koyanagi KO, Hagiwara M, Itoh T, Gojobori T, Imanishi T (July 2005). "Comparative genomics of bidirectional gene pairs and its implications for the evolution of a transcriptional regulation system". Gene. 353 (2): 169–176. doi:10.1016/j.gene.2005.04.027. PMID 15944140.
- ^ Liu B, Chen J, Shen B (May 2011). "Genome-wide analysis of the transcription factor binding preference of human bi-directional promoters and functional annotation of related gene pairs". BMC Systems Biology. 5 (Suppl 1): S2. doi:10.1186/1752-0509-5-S1-S2. PMC 3121118. PMID 21689477.
- ^ a b Shu J, Jelinek J, Chang H, Shen L, Qin T, Chung W, et al. (May 2006). "Silencing of bidirectional promoters by DNA methylation in tumorigenesis". Cancer Research. 66 (10): 5077–5084. doi:10.1158/0008-5472.CAN-05-2629. PMID 16707430.
- ^ a b Wei W, Pelechano V, Järvelin AI, Steinmetz LM (July 2011). "Functional consequences of bidirectional promoters". Trends in Genetics. 27 (7): 267–276. doi:10.1016/j.tig.2011.04.002. PMC 3123404. PMID 21601935.
- ^ Mahpour A, Scruggs BS, Smiraglia D, Ouchi T, Gelman IH (2018-10-17). "A methyl-sensitive element induces bidirectional transcription in TATA-less CpG island-associated promoters". PLOS ONE. 13 (10): e0205608. Bibcode:2018PLoSO..1305608M. doi:10.1371/journal.pone.0205608. PMC 6192621. PMID 30332484.
- ^ Lin JM, Collins PJ, Trinklein ND, Fu Y, Xi H, Myers RM, Weng Z (June 2007). "Transcription factor binding and modified histones in human bidirectional promoters". Genome Research. 17 (6): 818–827. doi:10.1101/gr.5623407. PMC 1891341. PMID 17568000.
- ^ Koev G, Miller WA (July 2000). "A positive-strand RNA virus with three very different subgenomic RNA promoters". Journal of Virology. 74 (13): 5988–5996. doi:10.1128/jvi.74.13.5988-5996.2000. PMC 112095. PMID 10846080.
- ^ deHaseth PL, Zupancic ML, Record MT (June 1998). "RNA polymerase-promoter interactions: the comings and goings of RNA polymerase". Journal of Bacteriology. 180 (12): 3019–3025. doi:10.1128/jb.180.12.3019-3025.1998. PMC 107799. PMID 9620948.
- ^ Singer P, Wu CW (October 1987). "Promoter search by Escherichia coli RNA polymerase on a circular DNA template". The Journal of Biological Chemistry. 262 (29): 14178–14189. doi:10.1016/S0021-9258(18)47921-5. PMID 3308887.
- ^ Borukhov S, Nudler E (April 2003). "RNA polymerase holoenzyme: structure, function and biological implications". Current Opinion in Microbiology. 6 (2): 93–100. doi:10.1016/s1369-5274(03)00036-5. PMID 12732296.
- ^ Copland JA, Sheffield-Moore M, Koldzic-Zivanovic N, Gentry S, Lamprou G, Tzortzatou-Stathopoulou F, et al. (June 2009). "Sex steroid receptors in skeletal differentiation and epithelial neoplasia: is tissue-specific intervention possible?". BioEssays. 31 (6): 629–641. doi:10.1002/bies.200800138. PMID 19382224. S2CID 205469320.
- ^ Vlahopoulos SA, Logotheti S, Mikas D, Giarika A, Gorgoulis V, Zoumpourlis V (April 2008). "The role of ATF-2 in oncogenesis". BioEssays. 30 (4): 314–327. doi:10.1002/bies.20734. PMID 18348191. S2CID 678541.
- ^ Saxonov S, Berg P, Brutlag DL (January 2006). "A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters". Proceedings of the National Academy of Sciences of the United States of America. 103 (5): 1412–1417. Bibcode:2006PNAS..103.1412S. doi:10.1073/pnas.0510310103. PMC 1345710. PMID 16432200.
- ^ Deaton AM, Bird A (May 2011). "CpG islands and the regulation of transcription". Genes & Development. 25 (10): 1010–1022. doi:10.1101/gad.2037511. PMC 3093116. PMID 21576262.
- ^ Chen HY, Shao CJ, Chen FR, Kwan AL, Chen ZP (April 2010). "Role of ERCC1 promoter hypermethylation in drug resistance to cisplatin in human gliomas". International Journal of Cancer. 126 (8): 1944–1954. doi:10.1002/ijc.24772. PMID 19626585.
- ^ Hossain A, Lopez E, Halper SM, Cetnar DP, Reis AC, Strickland D, et al. (December 2020). "Automated design of thousands of nonrepetitive parts for engineering stable genetic systems". Nature Biotechnology. 38 (12): 1466–1475. doi:10.1038/s41587-020-0584-2. PMID 32661437. S2CID 220506228.
- ^ Hobbs K, Negri J, Klinnert M, Rosenwasser LJ, Borish L (December 1998). "Interleukin-10 and transforming growth factor-beta promoter polymorphisms in allergies and asthma". American Journal of Respiratory and Critical Care Medicine. 158 (6): 1958–1962. doi:10.1164/ajrccm.158.6.9804011. PMID 9847292.
- ^ Burchard EG, Silverman EK, Rosenwasser LJ, Borish L, Yandava C, Pillari A, et al. (September 1999). "Association between a sequence variant in the IL-4 gene promoter and FEV(1) in asthma". American Journal of Respiratory and Critical Care Medicine. 160 (3): 919–922. doi:10.1164/ajrccm.160.3.9812024. PMID 10471619.
- ^ Kulozik AE, Bellan-Koch A, Bail S, Kohne E, Kleihauer E (May 1991). "Thalassemia intermedia: moderate reduction of beta globin gene transcriptional activity by a novel mutation of the proximal CACCC promoter element". Blood. 77 (9): 2054–2058. doi:10.1182/blood.V77.9.2054.2054. PMID 2018842.
- ^ Petrij F, Giles RH, Dauwerse HG, Saris JJ, Hennekam RC, Masuno M, et al. (July 1995). "Rubinstein-Taybi syndrome caused by mutations in the transcriptional co-activator CBP". Nature. 376 (6538): 348–351. Bibcode:1995Natur.376..348P. doi:10.1038/376348a0. PMID 7630403. S2CID 4254507.
- ^ Maloy S. "Expression vectors". San Diego State University.
External links
edit- ORegAnno – Open Regulatory Annotation Database
- Identifying a Protein Binding Sites on DNA molecule YouTube tutorial video
- Pleiades Promoter Project – a research project with an aim to generate 160 fully characterized, human DNA promoters of less than 4 kb (MiniPromoters) to drive gene expression in defined brain regions of therapeutic interests.
- ENCODE threads Explorer RNA and chromatin modification patterns around promoters. Nature (journal)