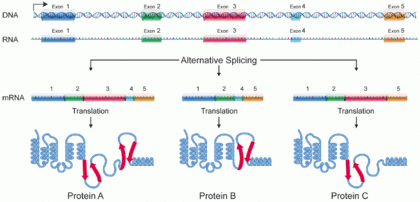
A protein isoform, or "protein variant",[1] is a member of a set of highly similar proteins that originate from a single gene and are the result of genetic differences.[2] While many perform the same or similar biological roles, some isoforms have unique functions. A set of protein isoforms may be formed from alternative splicings, variable promoter usage, or other post-transcriptional modifications of a single gene; post-translational modifications are generally not considered. (For that, see Proteoforms.) Through RNA splicing mechanisms, mRNA has the ability to select different protein-coding segments (exons) of a gene, or even different parts of exons from RNA to form different mRNA sequences. Each unique sequence produces a specific form of a protein.
The discovery of isoforms could explain the discrepancy between the small number of protein coding regions of genes revealed by the human genome project and the large diversity of proteins seen in an organism: different proteins encoded by the same gene could increase the diversity of the proteome. Isoforms at the RNA level are readily characterized by cDNA transcript studies. Many human genes possess confirmed alternative splicing isoforms. It has been estimated that ~100,000 expressed sequence tags (ESTs) can be identified in humans.[1] Isoforms at the protein level can manifest in the deletion of whole domains or shorter loops, usually located on the surface of the protein.[3]
Definition
editOne single gene has the ability to produce multiple proteins that differ both in structure and composition;[4][5] this process is regulated by the alternative splicing of mRNA, though it is not clear to what extent such a process affects the diversity of the human proteome, as the abundance of mRNA transcript isoforms does not necessarily correlate with the abundance of protein isoforms.[6] Three-dimensional protein structure comparisons can be used to help determine which, if any, isoforms represent functional protein products, and the structure of most isoforms in the human proteome has been predicted by AlphaFold and publicly released at isoform.io. [7] The specificity of translated isoforms is derived by the protein's structure/function, as well as the cell type and developmental stage during which they are produced.[4][5] Determining specificity becomes more complicated when a protein has multiple subunits and each subunit has multiple isoforms.
For example, the 5' AMP-activated protein kinase (AMPK), an enzyme, which performs different roles in human cells, has 3 subunits:[8]
- α, catalytic domain, has two isoforms: α1 and α2 which are encoded from PRKAA1 and PRKAA2
- β, regulatory domain, has two isoforms: β1 and β2 which are encoded from PRKAB1 and PRKAB2
- γ, regulatory domain, has three isoforms: γ1, γ2, and γ3 which are encoded from PRKAG1, PRKAG2, and PRKAG3
In human skeletal muscle, the preferred form is α2β2γ1.[8] But in the human liver, the most abundant form is α1β2γ1.[8]
Mechanism
editThe primary mechanisms that produce protein isoforms are alternative splicing and variable promoter usage, though modifications due to genetic changes, such as mutations and polymorphisms are sometimes also considered distinct isoforms.[9]
Alternative splicing is the main post-transcriptional modification process that produces mRNA transcript isoforms, and is a major molecular mechanism that may contribute to protein diversity.[5] The spliceosome, a large ribonucleoprotein, is the molecular machine inside the nucleus responsible for RNA cleavage and ligation, removing non-protein coding segments (introns).[10]
Because splicing is a process that occurs between transcription and translation, its primary effects have mainly been studied through genomics techniques—for example, microarray analyses and RNA sequencing have been used to identify alternatively spliced transcripts and measure their abundances.[9] Transcript abundance is often used as a proxy for the abundance of protein isoforms, though proteomics experiments using gel electrophoresis and mass spectrometry have demonstrated that the correlation between transcript and protein counts is often low, and that one protein isoform is usually dominant.[11] One 2015 study states that the cause of this discrepancy likely occurs after translation, though the mechanism is essentially unknown.[12] Consequently, although alternative splicing has been implicated as an important link between variation and disease, there is no conclusive evidence that it acts primarily by producing novel protein isoforms.[11]
Alternative splicing generally describes a tightly regulated process in which alternative transcripts are intentionally generated by the splicing machinery. However, such transcripts are also produced by splicing errors in a process called "noisy splicing," and are also potentially translated into protein isoforms. Although ~95% of multi-exonic genes are thought to be alternatively spliced, one study on noisy splicing observed that most of the different low-abundance transcripts are noise, and predicts that most alternative transcript and protein isoforms present in a cell are not functionally relevant.[13]
Other transcriptional and post-transcriptional regulatory steps can also produce different protein isoforms.[14] Variable promoter usage occurs when the transcriptional machinery of a cell (RNA polymerase, transcription factors, and other enzymes) begin transcription at different promoters—the region of DNA near a gene that serves as an initial binding site—resulting in slightly modified transcripts and protein isoforms.
Characteristics
editGenerally, one protein isoform is labeled as the canonical sequence based on criteria such as its prevalence and similarity to orthologous—or functionally analogous—sequences in other species.[15] Isoforms are assumed to have similar functional properties, as most have similar sequences, and share some to most exons with the canonical sequence. However, some isoforms show much greater divergence (for example, through trans-splicing), and can share few to no exons with the canonical sequence. In addition, they can have different biological effects—for example, in an extreme case, the function of one isoform can promote cell survival, while another promotes cell death—or can have similar basic functions but differ in their sub-cellular localization.[16] A 2016 study, however, functionally characterized all the isoforms of 1,492 genes and determined that most isoforms behave as "functional alloforms." The authors came to the conclusion that isoforms behave like distinct proteins after observing that the functional of most isoforms did not overlap.[17] Because the study was conducted on cells in vitro, it is not known if the isoforms in the expressed human proteome share these characteristics. Additionally, because the function of each isoform must generally be determined separately, most identified and predicted isoforms still have unknown functions.
Related concepts
editGlycoform
editA glycoform is an isoform of a protein that differs only with respect to the number or type of attached glycan. Glycoproteins often consist of a number of different glycoforms, with alterations in the attached saccharide or oligosaccharide. These modifications may result from differences in biosynthesis during the process of glycosylation, or due to the action of glycosidases or glycosyltransferases. Glycoforms may be detected through detailed chemical analysis of separated glycoforms, but more conveniently detected through differential reaction with lectins, as in lectin affinity chromatography and lectin affinity electrophoresis. Typical examples of glycoproteins consisting of glycoforms are the blood proteins as orosomucoid, antitrypsin, and haptoglobin. An unusual glycoform variation is seen in neuronal cell adhesion molecule, NCAM involving polysialic acids, PSA.
Examples
edit- G-actin: despite its conserved nature, it has a varying number of isoforms (at least six in mammals).
- Creatine kinase, the presence of which in the blood can be used as an aid in the diagnosis of myocardial infarction, exists in 3 isoforms.
- Hyaluronan synthase, the enzyme responsible for the production of hyaluronan, has three isoforms in mammalian cells.
- UDP-glucuronosyltransferase, an enzyme superfamily responsible for the detoxification pathway of many drugs, environmental pollutants, and toxic endogenous compounds has 16 known isoforms encoded in the human genome.[18]
- G6PDA: normal ratio of active isoforms in cells of any tissue is 1:1 shared with G6PDG. This is precisely the normal isoform ratio in hyperplasia. Only one of these isoforms is found during neoplasia.[19]
Monoamine oxidase, a family of enzymes that catalyze the oxidation of monoamines, exists in two isoforms, MAO-A and MAO-B.
See also
editReferences
edit- ^ a b Brett D, Pospisil H, Valcárcel J, Reich J, Bork P (January 2002). "Alternative splicing and genome complexity". Nature Genetics. 30 (1): 29–30. doi:10.1038/ng803. PMID 11743582. S2CID 2724843.
- ^ Schlüter H, Apweiler R, Holzhütter HG, Jungblut PR (September 2009). "Finding one's way in proteomics: a protein species nomenclature". Chemistry Central Journal. 3: 11. doi:10.1186/1752-153X-3-11. PMC 2758878. PMID 19740416.
- ^ Kozlowski, L.; Orlowski, J.; Bujnicki, J. M. (2012). "Structure Prediction for Alternatively Spliced Proteins". Alternative pre-mRNA Splicing. p. 582. doi:10.1002/9783527636778.ch54. ISBN 9783527636778.
- ^ a b Andreadis A, Gallego ME, Nadal-Ginard B (1987-01-01). "Generation of protein isoform diversity by alternative splicing: mechanistic and biological implications". Annual Review of Cell Biology. 3 (1): 207–42. doi:10.1146/annurev.cb.03.110187.001231. PMID 2891362.
- ^ a b c Breitbart RE, Andreadis A, Nadal-Ginard B (1987-01-01). "Alternative splicing: a ubiquitous mechanism for the generation of multiple protein isoforms from single genes". Annual Review of Biochemistry. 56 (1): 467–95. doi:10.1146/annurev.bi.56.070187.002343. PMID 3304142.
- ^ Liu Y, Beyer A, Aebersold R (April 2016). "On the Dependency of Cellular Protein Levels on mRNA Abundance". Cell. 165 (3): 535–50. doi:10.1016/j.cell.2016.03.014. hdl:20.500.11850/116226. PMID 27104977.
- ^ Sommer, Markus J.; Cha, Sooyoung; Varabyou, Ales; Rincon, Natalia; Park, Sukhwan; Minkin, Ilia; Pertea, Mihaela; Steinegger, Martin; Salzberg, Steven L. (2022-12-15). "Structure-guided isoform identification for the human transcriptome". eLife. 11: e82556. doi:10.7554/eLife.82556. PMC 9812405. PMID 36519529.
- ^ a b c Dasgupta B, Chhipa RR (March 2016). "Evolving Lessons on the Complex Role of AMPK in Normal Physiology and Cancer". Trends in Pharmacological Sciences. 37 (3): 192–206. doi:10.1016/j.tips.2015.11.007. PMC 4764394. PMID 26711141.
- ^ a b Kornblihtt AR, Schor IE, Alló M, Dujardin G, Petrillo E, Muñoz MJ (March 2013). "Alternative splicing: a pivotal step between eukaryotic transcription and translation". Nature Reviews Molecular Cell Biology. 14 (3): 153–65. doi:10.1038/nrm3525. hdl:11336/21049. PMID 23385723. S2CID 54560052.
- ^ Lee Y, Rio DC (2015-01-01). "Mechanisms and Regulation of Alternative Pre-mRNA Splicing". Annual Review of Biochemistry. 84 (1): 291–323. doi:10.1146/annurev-biochem-060614-034316. PMC 4526142. PMID 25784052.
- ^ a b Tress ML, Abascal F, Valencia A (February 2017). "Alternative Splicing May Not Be the Key to Proteome Complexity". Trends in Biochemical Sciences. 42 (2): 98–110. doi:10.1016/j.tibs.2016.08.008. PMC 6526280. PMID 27712956.
- ^ Battle A, Khan Z, Wang SH, Mitrano A, Ford MJ, Pritchard JK, Gilad Y (February 2015). "Genomic variation. Impact of regulatory variation from RNA to protein". Science. 347 (6222): 664–7. doi:10.1126/science.1260793. PMC 4507520. PMID 25657249.
- ^ Pickrell JK, Pai AA, Gilad Y, Pritchard JK (December 2010). "Noisy splicing drives mRNA isoform diversity in human cells". PLOS Genetics. 6 (12): e1001236. doi:10.1371/journal.pgen.1001236. PMC 3000347. PMID 21151575.
- ^ Smith LM, Kelleher NL (March 2013). "Proteoform: a single term describing protein complexity". Nature Methods. 10 (3): 186–7. doi:10.1038/nmeth.2369. PMC 4114032. PMID 23443629.
- ^ Li HD, Menon R, Omenn GS, Guan Y (December 2014). "Revisiting the identification of canonical splice isoforms through integration of functional genomics and proteomics evidence" (PDF). Proteomics. 14 (23–24): 2709–18. doi:10.1002/pmic.201400170. PMC 4372202. PMID 25265570.
- ^ Sundvall M, Veikkolainen V, Kurppa K, Salah Z, Tvorogov D, van Zoelen EJ, Aqeilan R, Elenius K (December 2010). "Cell death or survival promoted by alternative isoforms of ErbB4". Molecular Biology of the Cell. 21 (23): 4275–86. doi:10.1091/mbc.E10-04-0332. PMC 2993754. PMID 20943952.
- ^ Yang X, Coulombe-Huntington J, Kang S, Sheynkman GM, Hao T, Richardson A, et al. (February 2016). "Widespread Expansion of Protein Interaction Capabilities by Alternative Splicing". Cell. 164 (4): 805–17. doi:10.1016/j.cell.2016.01.029. PMC 4882190. PMID 26871637.
- ^ Barre L, Fournel-Gigleux S, Finel M, Netter P, Magdalou J, Ouzzine M (March 2007). "Substrate specificity of the human UDP-glucuronosyltransferase UGT2B4 and UGT2B7. Identification of a critical aromatic amino acid residue at position 33". The FEBS Journal. 274 (5): 1256–64. doi:10.1111/j.1742-4658.2007.05670.x. PMID 17263731.
- ^ Pathoma, Fundamentals of Pathology