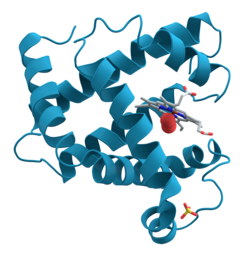
Ribbon diagrams, also known as Richardson diagrams, are 3D schematic representations of protein structure and are one of the most common methods of protein depiction used today. The ribbon depicts the general course and organisation of the protein backbone in 3D and serves as a visual framework for hanging details of the entire atomic structure, such as the balls for the oxygen atoms attached to myoglobin's active site in the adjacent figure. Ribbon diagrams are generated by interpolating a smooth curve through the polypeptide backbone. α-helices are shown as coiled ribbons or thick tubes, β-sheets as arrows, and non-repetitive coils or loops as lines or thin tubes. The direction of the polypeptide chain is shown locally by the arrows, and may be indicated overall by a colour ramp along the length of the ribbon.[1]
Ribbon diagrams are simple yet powerful, expressing the visual basics of a molecular structure (twist, fold and unfold). This method has successfully portrayed the overall organization of protein structures, reflecting their three-dimensional nature and allowing better understanding of these complex objects both by expert structural biologists and by other scientists, students,[2] and the general public.

History
editThe first ribbon diagrams, hand-drawn by Jane S. Richardson in 1980 (influenced by earlier individual illustrations),[3] were the first schematics of 3D protein structure to be produced systematically.[3][4] They were created to illustrate a classification of protein structures for an article in Advances in Protein Chemistry[5] (now available in annotated form on-line at Anatax). These drawings were outlined in pen on tracing paper over a printout of a Cα trace of the atomic coordinates, and shaded with colored pencil or pastels;[6] they preserved positions, smoothed the backbone path, and incorporated small local shifts to disambiguate the visual appearance.[4] As well as the triose isomerase ribbon drawing at the right, other hand-drawn examples depicted prealbumin, flavodoxin, and Cu,Zn superoxide dismutase.
In 1982, Arthur M. Lesk and co-workers first enabled the automatic generation of ribbon diagrams through a computational implementation that uses Protein Data Bank files as input.[7] This conceptually simple algorithm fit cubic polynomial B-spline curves to the peptide planes. Most modern graphics systems provide either B-splines or Hermite splines as a basic drawing primitive. One type of spline implementation passes through each Cα guide point, producing an exact but choppy curve. Both hand-drawn and most computer ribbons (such as those shown here) are smoothed over about four successive guide points (usually the peptide midpoint) to produce a more visually pleasing and understandable representation. To give the right radius for helical spirals while preserving smooth β-strands, the splines can be modified by offsets proportional to local curvature, as first developed by Mike Carson for his Ribbons program[8] and later adopted by other molecular graphics software, such as the open-source Mage program for kinemage graphics[9] that produced the ribbon image at top right (other examples: 1XK8 trimer and DNA polymerase).
Since their inception, and continuing in the present, ribbon diagrams have been the single most common representation of protein structure and a common choice of cover image for a journal or textbook.
Current computer programs
editOne popular program used for drawing ribbon diagrams is Molscript. Molscript utilizes Hermite splines to create coordinates for coils, turns, strands, and helices. The curve passes through all its control points (Cα atoms) guided by direction vectors. The program was built based on traditional molecular graphics by Arthur M. Lesk, Karl Hardman, and John Priestle.[10] Jmol is an open-source Java-based viewer for browsing molecular structures on the web; it includes a simplified "cartoon" version of ribbons. Other graphics programs such as DeepView (example: urease) and MolMol (example: SH2 domain) also produce ribbon images. KiNG[11] is the Java-based successor to Mage (examples: α-hemolysin top view and side view).
UCSF Chimera is a powerful molecular modeling program that also includes visualizations such as ribbons, notable especially for the ability to combine them with contoured shapes from cryo-electron microscopy data.[12] PyMOL, by Warren DeLano,[13] is a popular and flexible molecular graphics program (based on Python) that operates in interactive mode and also produces presentation-quality 2D images for ribbon diagrams and many other representations.
Features
edit| Secondary structure[4][5] | |
|---|---|
| α-Helices | Cylindrical spiral ribbons, with ribbon plane approximately following plane of peptides. |
| β-Strands | Arrows with thickness, about one-quarter as thick as they are wide, showing direction and twist of the strand from amino to carboxy end. β-sheets are seen as unified because neighboring strands twist in unison. |
| Loops and miscellaneous | |
| Nonrepetitive loops | Round ropes that are fatter in the foreground and thinner towards the back, following smoothed path of Cα trace. |
| Junctions between loops and helices | Round rope that gradually flattens out into a thin helical ribbon. |
| Other features | |
| Polypeptide direction,
NH2 and COOH termini |
Small arrows on one or both of the termini, or letters. For β-strands, the direction of the arrow is sufficient. Today, the direction of the polypeptide chain is often indicated by a colour ramp. |
| Disulfide bonds | Interlocked SS symbol or a zigzag, like a stylized lightning stroke. |
| Prosthetic groups or inhibitors | Stick figures, or ball & stick. |
| Metals | Spheres. |
| Shading and colour | Shading or colour adds dimensionality to the diagram. Generally, the features at the front are the highest in contrast and those towards the back are the lowest. |
See also
editReferences
edit- ^ Smith, Thomas J. (October 27, 2005). "Displaying and Analyzing Atomic Structures on the Macintosh". Danforth Plant Science Center. Archived from the original on 28 March 2002.
- ^ Richardson, D. C.; Richardson, J. S. (January 2002). "Teaching Molecular 3-D Literacy". Biochemistry and Molecular Biology Education. 30 (1): 21–26. doi:10.1002/bmb.2002.494030010005.
- ^ a b Richardson, Jane S. (2000), "Early ribbon drawings of proteins", Nature Structural Biology, 7 (8): 624–625, doi:10.1038/77912, PMID 10932243, S2CID 52856546.
- ^ a b c Richardson, Jane S. (1985), "Schematic drawings of protein structures", Diffraction Methods for Biological Macromolecules Part B, Methods in Enzymology, vol. 115, pp. 359–380, doi:10.1016/0076-6879(85)15026-3, ISBN 978-0-12-182015-2, PMID 3853075.
- ^ a b Richardson, Jane S. (1981), "The Anatomy and Taxonomy of Protein Structure", Anatomy and Taxonomy of Protein Structures, Advances in Protein Chemistry, vol. 34, pp. 167–339, doi:10.1016/S0065-3233(08)60520-3, ISBN 978-0-12-034234-1, PMID 7020376.
- ^ "Science's 'Mother of Ribbon Diagrams' celebrates 50 years at Duke". Duke Stories. 2018-10-19. Retrieved 2020-06-09.
- ^ Lesk, Arthur M.; Hardman, Karl D. (1982), "Computer-Generated Schematic Diagrams of Protein Structures", Science, 216 (4545): 539–540, Bibcode:1982Sci...216..539L, doi:10.1126/science.7071602, PMID 7071602.
- ^ Carson, M.; Bugg, C. E. (1986), "Algorithm for Ribbon Models of Proteins", Journal of Molecular Graphics, 4 (2): 121–122, doi:10.1016/0263-7855(86)80010-8.
- ^ Richardson, D. C.; Richardson, J. S. (January 1992), "The kinemage: a tool for scientific communication", Protein Science, 1 (1): 3–9, doi:10.1002/pro.5560010102, PMC 2142077, PMID 1304880
- ^ MolScript v2.1: About the program
- ^ Chen, V. B.; Davis, I. W.; Richardson, D. C. (2009), "KING (Kinemage, Next Generation): A versatile interactive molecular and scientific visualization program", Protein Science, 18 (11): 2403–2409, doi:10.1002/pro.250, PMC 2788294, PMID 19768809
- ^ Goddard, Thomas D.; Huang, Conrad C.; Ferrin, Thomas E. (2005), "Software Extensions to UCSF Chimera for Interactive Visualization of Large Molecular Assemblies", Structure, 13 (3): 473–482, doi:10.1016/j.str.2005.01.006, PMID 15766548.
- ^ Brunger, Axel T.; Wells, James A. (2009), "Warren L. DeLano, 21 June 1972-3 November 2009", Nature Structural & Molecular Biology, 16 (12): 1202–1203, doi:10.1038/nsmb1209-1202, PMID 19956203.