The RprA RNA gene encodes a 106 nucleotide regulatory non-coding RNA. Translational regulation of the stationary phase sigma factor RpoS is mediated by the formation of a double-stranded RNA stem-loop structure in the upstream region of the rpoS messenger RNA, occluding the translation initiation site.[1][2]
| RprA RNA | |
|---|---|
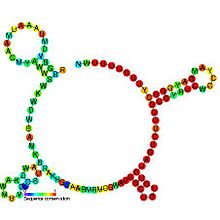 Predicted secondary structure and sequence conservation of RprA | |
| Identifiers | |
| Symbol | RprA |
| Rfam | RF00034 |
| Other data | |
| RNA type | Gene; sRNA |
| Domain(s) | Bacteria |
| SO | SO:0000387 |
| PDB structures | PDBe |
Clones carrying rprA (RpoS regulator RNA A) increased the translation of RpoS. As with DsrA, RprA is predicted to form three stem-loops. Thus, at least two small RNAs, DsrA and RprA, participate in the positive regulation of RpoS translation. RprA also appears to bind to the RpoS leader.[3] RprA is non-essential.[4] Wasserman et al. demonstrated that this RNA is bound by the Hfq protein.[5] Binding to Hfq alters the conformation of RprA.[6] In the presence of Hfq the stability of RprA is influenced by the osmolarity of the cell, this is dependent on the endoribonuclease RNase E.[7]
It has been shown the RprA regulates the protein coding genes, called csgD, this protein encodes a stationary phase-induced biofilm regulator and ydaM, which encodes a diguanylate cyclase involved in activating csgD transcription. These two target genes are repressed by RprA which results in regulation of biofilm formation.[8]
References
edit- ^ Updegrove T, Wilf N, Sun X, Wartell RM (2008). "Effect of Hfq on RprA-rpoS mRNA pairing: Hfq-RNA binding and the influence of the 5′ rpoS mRNA leader region". Biochemistry. 47 (43): 11184–11195. doi:10.1021/bi800479p. PMID 18826256.
- ^ Jones AM, Goodwill A, Elliott T (2006). "Limited Role for the DsrA and RprA Regulatory RNAs in rpoS Regulation in Salmonella enterica". J Bacteriol. 188 (14): 5077–5088. doi:10.1128/JB.00206-06. PMC 1539969. PMID 16816180.
- ^ Majdalani, N; Hernandez D; Gottesman S (2002). "Regulation and mode of action of the second small RNA activator of RpoS translation, RprA". Mol Microbiol. 46 (3): 813–826. doi:10.1046/j.1365-2958.2002.03203.x. PMID 12410838.
- ^ Majdalani, N; Chen S; Murrow J; St John K; Gottesman S (2001). "Regulation of RpoS by a novel small RNA: the characterization of RprA". Mol Microbiol. 39 (5): 1382–1394. doi:10.1111/j.1365-2958.2001.02329.x. PMID 11251852. S2CID 90988055.
- ^ Wassarman KM, Repoila F, Rosenow C, Storz G, Gottesman S (2001). "Identification of novel small RNAs using comparative genomics and microarrays". Genes Dev. 15 (13): 1637–1651. doi:10.1101/gad.901001. PMC 312727. PMID 11445539.
- ^ Henderson CA, Vincent HA, Casamento A, Stone CM, Phillips JO, Cary PD, Sobott F, Gowers DM, Taylor JE, Callaghan AJ (August 2013). "Hfq binding changes the structure of Escherichia coli small noncoding RNAs OxyS and RprA, which are involved in the riboregulation of rpoS". RNA. 19 (8): 1089–1104. doi:10.1261/rna.034595.112. PMC 3708529. PMID 23804244.
- ^ Madhugiri R, Basineni SR, Klug G (2010). "Turn-over of the small non-coding RNA RprA in E. coli is influenced by osmolarity". Mol Genet Genomics. 284 (4): 307–318. doi:10.1007/s00438-010-0568-x. PMID 20717695. S2CID 23732166.
- ^ Mika F, Busse S, Possling A, Berkholz J, Tschowri N, Sommerfeldt N, Pruteanu M, Hengge R (2012). "Targeting of csgD by the small regulatory RNA RprA links stationary phase, biofilm formation and cell envelope stress in Escherichia coli". Mol Microbiol. 84 (1): 51–65. doi:10.1111/j.1365-2958.2012.08002.x. PMC 3465796. PMID 22356413.