Chromosome 3 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 3 spans more than 198 million base pairs (the building material of DNA) and represents about 6.5 percent of the total DNA in cells.
| Chromosome 3 | |
|---|---|
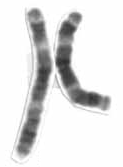 Human chromosome 3 pair after G-banding. One is from mother, one is from father. | |
 Chromosome 3 pair in human male karyogram. | |
| Features | |
| Length (bp) | 201,105,948 bp (CHM13) |
| No. of genes | 1,024 (CCDS)[1] |
| Type | Autosome |
| Centromere position | Metacentric[2] (90.9 Mbp[3]) |
| Complete gene lists | |
| CCDS | Gene list |
| HGNC | Gene list |
| UniProt | Gene list |
| NCBI | Gene list |
| External map viewers | |
| Ensembl | Chromosome 3 |
| Entrez | Chromosome 3 |
| NCBI | Chromosome 3 |
| UCSC | Chromosome 3 |
| Full DNA sequences | |
| RefSeq | NC_000003 (FASTA) |
| GenBank | CM000665 (FASTA) |
Genes
editNumber of genes
editThe following are some of the gene count estimates of human chromosome 3. Because researchers use different approaches to genome annotation their predictions of the number of genes on each chromosome varies (for technical details, see gene prediction). Among various projects, the collaborative consensus coding sequence project (CCDS) takes an extremely conservative strategy. So CCDS's gene number prediction represents a lower bound on the total number of human protein-coding genes.[4]
| Estimated by | Protein-coding genes | Non-coding RNA genes | Pseudogenes | Source | Release date |
|---|---|---|---|---|---|
| CCDS | 1,024 | — | — | [1] | 2016-09-08 |
| HGNC | 1,036 | 483 | 761 | [5] | 2017-05-12 |
| Ensembl | 1,073 | 1,158 | 761 | [6] | 2017-03-29 |
| UniProt | 1,081 | — | — | [7] | 2018-02-28 |
| NCBI | 1,085 | 1,108 | 902 | [8][9][10] | 2017-05-19 |
List of genes
editThe following is a partial list of genes on human chromosome 3. For complete list, see the link in the infobox on the right.
p-arm
editPartial list of the genes located on p-arm (short arm) of human chromosome 3:
- ALAS1: aminolevulinate, delta-, synthase 1
- APEH: encoding enzyme Acylamino-acid-releasing enzyme
- ARPP-21: Cyclic AMP-regulated phosphoprotein, 21 kDa
- AZI2: encoding protein 5-azacytidine-induced protein 2
- BRK1: SCAR/WAVE actin nucleating complex subunit
- BRPF1: bromodomain and PHD finger containing 1
- BTD: biotinidase
- C3orf14-Chromosome 3 open reading frame 14: predicted DNA binding protein.
- CFAP20DC: encoding protein Chromosome 3 open reading frame 67
- C3orf62: chromosome 3 open reading frame 62
- CACNA2D3: calcium channel, voltage-dependent, alpha 2/delta subunit 3
- CCR5: chemokine (C-C motif) receptor 5
- CGGBP1: CGG triplet repeat binding protein 1
- CMTM7: CKLF like MARVEL transmembrane domain containing 7
- CNTN4: Contactin 4
- COL7A1: Collagen, type VII, alpha 1 (epidermolysis bullosa, dystrophic, dominant and recessive)
- CRBN: Cereblon protein[11]
- DCLK3: Doublecortin like kinase 3
- DLEC1: encoding protein Deleted in lung and esophageal cancer 1
- EAF1: ELL associated factor 1
- ENTPD3: ectonucleoside triphosphate diphosphohydrolase 3
- FAM107A: Family with sequence similarity 107 member A
- FAM19A1: Family with sequence similarity 19 member A1, C-C motif chemokine like
- FBXL2: F-box and leucine rich repeat protein 2
- FOXP1: Forkhead Box Protein P1
- FRA3A encoding protein Fragile site, aphidicolin type, common, fra(3)(p24.2)
- FRMD4B encoding protein FERM domain containing 4B
- GHRLOS: non-coding RNA ghrelin opposite strand (non-protein coding)
- GMPPB: GDP-mannose pyrophosphorylase B
- HACL1: encoding protein 2-hydroxyacyl-CoA lyase 1
- HEMK1: encoding protein HemK methyltransferase family member 1
- HIGD1A: HIG1 domain family member 1A
- HSN1B: Hereditary sensory neuropathy, type ib
- HTD2: encoding protein Hydroxyacyl-thioester dehydratase type 2
- LARS2: leucyl-tRNA synthetase, mitochondrial
- LIMD1: LIM domain-containing protein 1
- LOC105377021: encoding protein LOC105377021
- LINC00312: Long intergenic non-protein-coding RNA 312
- LZTFL1: Leucine zipper transcription factor like 1
- MIR138-1: encoding protein MicroRNA 138-1
- MIR425: MicroRNA 425
- MIR885: encoding protein MicroRNA 885
- MITF: microphthalmia-associated transcription factor
- MLH1: mutL homolog 1, colon cancer, nonpolyposis type 2 (E. coli)
- MYRIP: Myosin VIIA and Rab interacting protein
- NBEAL2: Neurobeachin-like 2
- NDUFAF3: encoding enzyme NADH dehydrogenase [ubiquinone] 1 alpha subcomplex assembly factor 3
- NKTR: NK-tumor recognition protein
- NPRL2: Nitrogen permease regulator 2-like protein
- OXTR: oxytocin receptor
- PCAF: acetyltransferase activity
- PHF7 encoding protein PHD finger protein 7
- PRICKLE2: encoding protein Prickle planar cell polarity protein 2
- PTHR1: parathyroid hormone receptor 1
- QRICH1: encoding protein QRICH1, also known as Glutamine-rich protein 1,
- RBM6: RNA-binding protein 6
- RPP14: Ribonuclease P protein subunit p14
- SCN5A: sodium channel, voltage-gated, type V, alpha (long QT syndrome 3)
- SETD5: SET domain containing 5
- SFMBT1: Scm-like with four mbt domains 1
- SLC25A20: solute carrier family 25 (carnitine/acylcarnitine translocase), member 20
- STT3B: catalytic subunit of the oligosaccharyltransferase complex
- SYNPR: synaptoporin
- TAFA4: encoding protein Family with sequence similarity 19 member A4, C-C motif chemokine like
- TCAIM: encoding protein T-cell activation inhibitor, mitochondrial
- TDGF1: Teratocarcinoma-derived growth factor 1
- TMEM158: Transmembrane protein 158
- TMIE: transmembrane inner ear
- TRAK1: trafficking kinesin-binding protein 1
- TRANK1: encoding protein Tetratricopeptide repeat and ankyrin repeat containing 1
- TTLL3: encoding protein Tubulin tyrosine ligase-like family, member 3
- TUSC2: tumor suppressor candidate 2
- UCN2: Urocortin-2
- ULK4: UNC-51 like kinase 4
- VGLL3: vestigial-like family member 3
- VHL: von Hippel-Lindau tumor suppressor
- ZMYND10: zinc finger MYND-type containing 10
- ZNF197: encoding protein Zinc finger protein 197
- ZNF502: encoding protein Zinc finger protein 502
- ZNF620: encoding protein Zinc finger protein 620
- ZNF621: encoding protein Zinc finger protein 621
- ZNF717: encoding protein Zinc finger protein 717
q-arm
editPartial list of the genes located on q-arm (long arm) of human chromosome 3:
- ADIPOQ: adiponectin
- AMOTL2: encoding protein Angiomotin-like protein 2
- ARHGAP31: Rho GRPase activating protein 31
- BCHE: butyrylcholinesterase
- C3orf70 chromosome 3 open reading frame 70
- CAMPD1: Camptodactyly
- CCDC80: Coiled-coil domain containing protein 80
- CD200R1: Cell surface glycoprotein CD200 receptor 1
- CHST13: encoding protein Carbohydrate (chondroitin 4) sulfotransferase 13
- CLDND1: Claudin domain containing 1
- CPN2: Carboxypeptidase N subunit 2
- CPOX: coproporphyrinogen oxidase (coproporphyria, harderoporphyria)
- DPPA2: Developmental pluripotency associated 2
- DTX3L: encoding protein Deltex e3 ubiquitin ligase 3l
- DZIP3: encoding protein DAZ interacting zinc finger protein 3
- EAF2: ELL associated factor 2
- EFCC1: EF-hand and coiled-coil domain containing 1
- ETM1: Essential tremor 1
- ETV5: ETS variant 5
- FAM3D: family with sequence similarity 3, member D
- FAM43A: family with sequence similarity 43 member A
- FAM162A: family with sequence similarity 162 member A
- FBXO40: encoding protein F-box protein 40
- FILIP1L: encoding protein Filamin A interacting protein 1 like
- GYG1: Glycogenin-1
- HACD2 encoding protein 3-hydroxyacyl-CoA dehydratase 2
- HGD: homogentisate 1,2-dioxygenase (homogentisate oxidase)
- IFT122: intraflagellar transport gene 122
- KIAA1257: KIAA1257
- LINC01279: encoding protein long intergenic non-protein coding RNA 1279
- LNCR5: encoding protein lung cancer susceptibility 5
- LMLN: encoding protein Leishmanolysin-like (metallopeptidase M8 family)
- LRRC15: leucine rich repeat containing 15
- LSG1: large subunit GTPase 1 homolog
- MB21D2: encoding protein Mab-21 domain containing 2
- MCCC1: methylcrotonoyl-Coenzyme A carboxylase 1 (alpha)
- MORC1: encoding protein Morc family cw-type zinc finger 1
- MYLK: Telokin
- NEPRO: encoding protein Nucleolus and neural progenitor protein
- NFKBIZ: NF-kappa-B inhibitor zeta
- OTOL1: encoding glycoprotein Otolin
- PARP14 encoding protein Poly(ADP-ribose) polymerase family member 14
- PCCB: propionyl Coenzyme A carboxylase, beta polypeptide
- PDCD10: programmed cell death 10
- PIK3CA: phosphoinositide-3-kinase, catalytic, alpha polypeptide
- PISRT1: long non-coding RNA
- PROSER1: Proline and serine rich protein 1
- RAB7: RAB7, member RAS oncogene family
- RASA2: encoding protein Ras p21 protein activator 2
- RETNLB: resistin-like beta
- RHO: rhodopsin visual pigment
- RIOX2: Ribosomal oxygenase 2
- SELT: Selenoprotein T
- SENP7: Sentrin-specific protease 7
- SERP1: Stress-associated endoplasmic reticulum protein 1
- SOX2: transcription factor
- SOX2OT: SOX2 overlapping transcript
- SPG14 encoding protein Spastic paraplegia 14 (autosomal recessive)
- SRPRB: Signal recognition particle receptor subunit beta
- TEX55: encoding protein Testis expressed 55
- TIMMDC1: TIMMDC1
- TMEM44: encoding protein Transmembrane protein 44
- TM4SF1: Transmembrane 4 L6 family member 1
- TMPRSS7: encoding protein Transmembrane serine protease 7
- TP63: Tumor protein p63
- TRAT1: T-cell receptor-associated transmembrane adapter 1
- USH3A: Usher syndrome 3A
- ZBED2: encoding protein Zinc finger BED-type containing 2
- ZNF9: zinc finger protein 9 (a cellular retroviral nucleic acid binding protein)
Diseases and disorders
editThe following diseases and disorders are some of those related to genes on chromosome 3:
- 3-Methylcrotonyl-CoA carboxylase deficiency
- 3q29 microdeletion syndrome
- Acute myeloid leukemia (AML)
- Alkaptonuria
- Arrhythmogenic right ventricular dysplasia
- Atransferrinemia
- Autism
- Autosomal dominant optic atrophy
- ADOA plus syndrome
- Biotinidase deficiency
- Blepharophimosis, epicanthus inversus and ptosis type 1
- Breast/colon/lung/pancreatic cancer
- Brugada syndrome
- Castillo fever
- Carnitine-acylcarnitine translocase deficiency
- Cataracts
- Cerebral cavernous malformation
- Charcot–Marie–Tooth disease, type 2
- Charcot–Marie–Tooth disease
- Chromosome 3q duplication syndrome
- Coproporphyria
- A location on human chromosome 3 is associated with respiratory failure and possibly with increased severity in COVID-19[12][13]
- Dandy–Walker syndrome
- Deafness
- Diabetes
- Dystrophic epidermolysis bullosa
- Endplate acetylcholinesterase deficiency
- Essential tremors
- Ectrodactyly, Case 4
- Glaucoma, primary open angle
- Glycogen storage disease
- Hailey–Hailey disease
- Harderoporphyrinuria
- Heart block, progressive/nonprogressive
- Hereditary coproporphyria
- Hereditary nonpolyposis colorectal cancer
- HIV infection, susceptibility/resistance to
- Hypobetalipoproteinemia, familial
- Hypothermia
- Leukoencephalopathy with vanishing white matter
- Long QT syndrome
- Lymphomas
- Malignant hyperthermia susceptibility
- Metaphyseal chondrodysplasia, Murk Jansen type
- Microcoria
- Möbius syndrome
- Moyamoya disease
- Mucopolysaccharidosis
- Muir–Torre family cancer syndrome
- Myotonic dystrophy
- Neuropathy, hereditary motor and sensory, Okinawa type
- Night blindness
- Nonsyndromic deafness
- Ovarian cancer
- Porphyria
- Propionic acidemia
- Protein S deficiency
- Pseudocholinesterase deficiency
- Pseudo-Zellweger syndrome
- Retinitis pigmentosa
- Romano–Ward syndrome
- Seckel syndrome
- Sensenbrenner syndrome
- Septo-optic dysplasia
- Short stature
- Spinocerebellar ataxia
- Sucrose intolerance
- T-cell leukemia translocation altered gene
- Usher syndrome
- von Hippel–Lindau syndrome
- Waardenburg syndrome
- Xeroderma pigmentosum, complementation group c
Cytogenetic band
edit| Chr. | Arm[18] | Band[19] | ISCN start[20] |
ISCN stop[20] |
Basepair start |
Basepair stop |
Stain[21] | Density |
|---|---|---|---|---|---|---|---|---|
| 3 | p | 26.3 | 0 | 175 | 1 | 2,800,000 | gpos | 50 |
| 3 | p | 26.2 | 175 | 263 | 2,800,001 | 4,000,000 | gneg | |
| 3 | p | 26.1 | 263 | 408 | 4,000,001 | 8,100,000 | gpos | 50 |
| 3 | p | 25.3 | 408 | 642 | 8,100,001 | 11,600,000 | gneg | |
| 3 | p | 25.2 | 642 | 759 | 11,600,001 | 13,200,000 | gpos | 25 |
| 3 | p | 25.1 | 759 | 963 | 13,200,001 | 16,300,000 | gneg | |
| 3 | p | 24.3 | 963 | 1269 | 16,300,001 | 23,800,000 | gpos | 100 |
| 3 | p | 24.2 | 1269 | 1357 | 23,800,001 | 26,300,000 | gneg | |
| 3 | p | 24.1 | 1357 | 1561 | 26,300,001 | 30,800,000 | gpos | 75 |
| 3 | p | 23 | 1561 | 1751 | 30,800,001 | 32,000,000 | gneg | |
| 3 | p | 22.3 | 1751 | 1926 | 32,000,001 | 36,400,000 | gpos | 50 |
| 3 | p | 22.2 | 1926 | 2013 | 36,400,001 | 39,300,000 | gneg | |
| 3 | p | 22.1 | 2013 | 2188 | 39,300,001 | 43,600,000 | gpos | 75 |
| 3 | p | 21.33 | 2188 | 2451 | 43,600,001 | 44,100,000 | gneg | |
| 3 | p | 21.32 | 2451 | 2626 | 44,100,001 | 44,200,000 | gpos | 50 |
| 3 | p | 21.31 | 2626 | 3239 | 44,200,001 | 50,600,000 | gneg | |
| 3 | p | 21.2 | 3239 | 3385 | 50,600,001 | 52,300,000 | gpos | 25 |
| 3 | p | 21.1 | 3385 | 3676 | 52,300,001 | 54,400,000 | gneg | |
| 3 | p | 14.3 | 3676 | 3910 | 54,400,001 | 58,600,000 | gpos | 50 |
| 3 | p | 14.2 | 3910 | 4143 | 58,600,001 | 63,800,000 | gneg | |
| 3 | p | 14.1 | 4143 | 4362 | 63,800,001 | 69,700,000 | gpos | 50 |
| 3 | p | 13 | 4362 | 4566 | 69,700,001 | 74,100,000 | gneg | |
| 3 | p | 12.3 | 4566 | 4814 | 74,100,001 | 79,800,000 | gpos | 75 |
| 3 | p | 12.2 | 4814 | 4946 | 79,800,001 | 83,500,000 | gneg | |
| 3 | p | 12.1 | 4946 | 5077 | 83,500,001 | 87,100,000 | gpos | 75 |
| 3 | p | 11.2 | 5077 | 5135 | 87,100,001 | 87,800,000 | gneg | |
| 3 | p | 11.1 | 5135 | 5266 | 87,800,001 | 90,900,000 | acen | |
| 3 | q | 11.1 | 5266 | 5427 | 90,900,001 | 94,000,000 | acen | |
| 3 | q | 11.2 | 5427 | 5602 | 94,000,001 | 98,600,000 | gvar | |
| 3 | q | 12.1 | 5602 | 5762 | 98,600,001 | 100,300,000 | gneg | |
| 3 | q | 12.2 | 5762 | 5850 | 100,300,001 | 101,200,000 | gpos | 25 |
| 3 | q | 12.3 | 5850 | 5996 | 101,200,001 | 103,100,000 | gneg | |
| 3 | q | 13.11 | 5996 | 6229 | 103,100,001 | 106,500,000 | gpos | 75 |
| 3 | q | 13.12 | 6229 | 6361 | 106,500,001 | 108,200,000 | gneg | |
| 3 | q | 13.13 | 6361 | 6594 | 108,200,001 | 111,600,000 | gpos | 50 |
| 3 | q | 13.2 | 6594 | 6682 | 111,600,001 | 113,700,000 | gneg | |
| 3 | q | 13.31 | 6682 | 6871 | 113,700,001 | 117,600,000 | gpos | 75 |
| 3 | q | 13.32 | 6871 | 6973 | 117,600,001 | 119,300,000 | gneg | |
| 3 | q | 13.33 | 6973 | 7148 | 119,300,001 | 122,200,000 | gpos | 75 |
| 3 | q | 21.1 | 7148 | 7294 | 122,200,001 | 124,100,000 | gneg | |
| 3 | q | 21.2 | 7294 | 7440 | 124,100,001 | 126,100,000 | gpos | 25 |
| 3 | q | 21.3 | 7440 | 7674 | 126,100,001 | 129,500,000 | gneg | |
| 3 | q | 22.1 | 7674 | 7936 | 129,500,001 | 134,000,000 | gpos | 25 |
| 3 | q | 22.2 | 7936 | 8053 | 134,000,001 | 136,000,000 | gneg | |
| 3 | q | 22.3 | 8053 | 8228 | 136,000,001 | 139,000,000 | gpos | 25 |
| 3 | q | 23 | 8228 | 8461 | 139,000,001 | 143,100,000 | gneg | |
| 3 | q | 24 | 8461 | 8811 | 143,100,001 | 149,200,000 | gpos | 100 |
| 3 | q | 25.1 | 8811 | 9001 | 149,200,001 | 152,300,000 | gneg | |
| 3 | q | 25.2 | 9001 | 9162 | 152,300,001 | 155,300,000 | gpos | 50 |
| 3 | q | 25.31 | 9162 | 9264 | 155,300,001 | 157,300,000 | gneg | |
| 3 | q | 25.32 | 9264 | 9366 | 157,300,001 | 159,300,000 | gpos | 50 |
| 3 | q | 25.33 | 9366 | 9453 | 159,300,001 | 161,000,000 | gneg | |
| 3 | q | 26.1 | 9453 | 9803 | 161,000,001 | 167,900,000 | gpos | 100 |
| 3 | q | 26.2 | 9803 | 9949 | 167,900,001 | 171,200,000 | gneg | |
| 3 | q | 26.31 | 9949 | 10183 | 171,200,001 | 176,000,000 | gpos | 75 |
| 3 | q | 26.32 | 10183 | 10329 | 176,000,001 | 179,300,000 | gneg | |
| 3 | q | 26.33 | 10329 | 10489 | 179,300,001 | 183,000,000 | gpos | 75 |
| 3 | q | 27.1 | 10489 | 10620 | 183,000,001 | 184,800,000 | gneg | |
| 3 | q | 27.2 | 10620 | 10737 | 184,800,001 | 186,300,000 | gpos | 25 |
| 3 | q | 27.3 | 10737 | 10883 | 186,300,001 | 188,200,000 | gneg | |
| 3 | q | 28 | 10883 | 11175 | 188,200,001 | 192,600,000 | gpos | 75 |
| 3 | q | 29 | 11175 | 11700 | 192,600,001 | 198,295,559 | gneg |
See also
editReferences
edit- ^ a b "Search results – 3[CHR] AND "Homo sapiens"[Organism] AND ("has ccds"[Properties] AND alive[prop]) – Gene". NCBI. CCDS Release 20 for Homo sapiens. 2016-09-08. Retrieved 2017-05-28.
- ^ Tom Strachan; Andrew Read (2 April 2010). Human Molecular Genetics. Garland Science. p. 45. ISBN 978-1-136-84407-2.
- ^ a b c Genome Decoration Page, NCBI. Ideogram data for Homo sapience (850 bphs, Assembly GRCh38.p3). Last update 2014-06-03. Retrieved 2017-04-26.
- ^ Pertea M, Salzberg SL (2010). "Between a chicken and a grape: estimating the number of human genes". Genome Biol. 11 (5): 206. doi:10.1186/gb-2010-11-5-206. PMC 2898077. PMID 20441615.
- ^ "Statistics & Downloads for chromosome 3". HUGO Gene Nomenclature Committee. 2017-05-12. Archived from the original on 2017-06-29. Retrieved 2017-05-19.
- ^ "Chromosome 3: Chromosome summary – Homo sapiens". Ensembl Release 88. 2017-03-29. Retrieved 2017-05-19.
- ^ "Human chromosome 3: entries, gene names and cross-references to MIM". UniProt. 2018-02-28. Retrieved 2018-03-16.
- ^ "Search results – 3[CHR] AND "Homo sapiens"[Organism] AND ("genetype protein coding"[Properties] AND alive[prop]) – Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- ^ "Search results – 3[CHR] AND "Homo sapiens"[Organism] AND ( ("genetype miscrna"[Properties] OR "genetype ncrna"[Properties] OR "genetype rrna"[Properties] OR "genetype trna"[Properties] OR "genetype scrna"[Properties] OR "genetype snrna"[Properties] OR "genetype snorna"[Properties]) NOT "genetype protein coding"[Properties] AND alive[prop]) – Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- ^ "Search results – 3[CHR] AND "Homo sapiens"[Organism] AND ("genetype pseudo"[Properties] AND alive[prop]) – Gene". NCBI. 2017-05-19. Retrieved 2017-05-20.
- ^ CRBN cereblon [Homo sapiens (human)] - Gene - NCBI
- ^ "Scientists pinpoint genes common among people with severe coronavirus infections". MSN.
- ^ Severe Covid-19 GWAS Group; et al. (2020). "Genomewide Association Study of Severe Covid-19 with Respiratory Failure". New England Journal of Medicine. 383 (16): 1522–1534. doi:10.1056/NEJMoa2020283. PMC 7315890. PMID 32558485.
{{cite journal}}: CS1 maint: numeric names: authors list (link) - ^ Genome Decoration Page, NCBI. Ideogram data for Homo sapience (400 bphs, Assembly GRCh38.p3). Last update 2014-03-04. Retrieved 2017-04-26.
- ^ Genome Decoration Page, NCBI. Ideogram data for Homo sapience (550 bphs, Assembly GRCh38.p3). Last update 2015-08-11. Retrieved 2017-04-26.
- ^ International Standing Committee on Human Cytogenetic Nomenclature (2013). ISCN 2013: An International System for Human Cytogenetic Nomenclature (2013). Karger Medical and Scientific Publishers. ISBN 978-3-318-02253-7.
- ^ Sethakulvichai, W.; Manitpornsut, S.; Wiboonrat, M.; Lilakiatsakun, W.; Assawamakin, A.; Tongsima, S. (2012). "Estimation of band level resolutions of human chromosome images". 2012 Ninth International Conference on Computer Science and Software Engineering (JCSSE). pp. 276–282. doi:10.1109/JCSSE.2012.6261965. ISBN 978-1-4673-1921-8. S2CID 16666470.
- ^ "p": Short arm; "q": Long arm.
- ^ For cytogenetic banding nomenclature, see article locus.
- ^ a b These values (ISCN start/stop) are based on the length of bands/ideograms from the ISCN book, An International System for Human Cytogenetic Nomenclature (2013). Arbitrary unit.
- ^ gpos: Region which is positively stained by G banding, generally AT-rich and gene poor; gneg: Region which is negatively stained by G banding, generally CG-rich and gene rich; acen Centromere. var: Variable region; stalk: Stalk.
External links
edit- National Institutes of Health. "Chromosome 3". Genetics Home Reference. Archived from the original on 2010-04-08. Retrieved 2017-05-06.
- "Chromosome 3". Human Genome Project Information Archive 1990–2003. Retrieved 2017-05-06.